▻ Overview
▻ Stage Description
▻ Images
▻ Download Supplemental Table
▻ In Situ Hybridization Data
▻ Sequences
Overview
A stage of Smed embryonic development defined by a unique gene expression signature and morphology, 9 - 11 days post-egg capsule deposition at 20C. Organogenesis and organ maturation continues. Embryos elongated. Eyes just visible. Onset of gliding motility. [PMID:28072387]
Stage Description
Elongated embryos continue to flatten, but have clearly adopted the bilaterally symmetric body plan of juvenile and adult animals. Pigmentation, if present, is sparse. Animals are motile: slow gliding locomotion is apparent, but animals cannot flip over. Mature, ciliated epidermal cells largely cover the dorsal and ventral surfaces of the embryos. A dense network of elongated, anteroposterior and circular fibers of body wall muscle is evident beneath the epidermis. Gut development continues: branching morphogenesis is evident, as is clear separation of the two posterior gut branches. piwi-1+ cells occupy the parenchymal space between the gut branches and along the tail stripe. Expression of embryonic gut markers ceases, and expression of mature gut markers becomes robust. Resorption of the temporary embryonic pharynx is complete, and definitive pharynx formation continues. The bilobed brain and ventral nerve cords are well developed, and lateral and commissural neurons continue to develop. The eyes, noted by the presence of developing pigment cups in live animals, first become visible during S7.
Images


Download Supplemental Table
Figure 1 – source data 7. S7 enriched transcripts from pairwise and/or mixed stage reference comparisons
Criteria for flagging differentially expressed transcripts:
Criteria for flagging differentially expressed transcripts: 1) Pairwise comparison (S6 vs S7): adjusted p-value < 1e-5 S7 vs mixed reference comparison: adjusted p-value <1e-20 2) log2 ratio ≥ 2.322 (5-fold upregulation, both comparisons) or log2 ratio ≤ -2.322 (5 fold downregulation, pairwise comparison only) 3) Average scaled RPKM value ≥ 1.0 for S7 4) Transcript has ≥ 1 ORF 5) Transcripts derived from transposase or retroviruses were omitted.
Tabs in this excel file contain 1) pairwise comparison data (if applicable), 2) mixed stage reference comparison data, 3) cluster membership, average RPKM values across embryogenesis (Y-S8), and in C4 and SX adults, as well as best BLASTx hits (E < 0.001) versus the NR, Swiss-Prot, C. elegans, D. melanogaster, D. rerio, X. tropicalis, M. musculus and H. sapiens REF-Seq databases, 4) GO analysis: Manually curated and categorized Biological Process (BP) GO IDs and 5) GO analysis: unabridged results. S2-S8: Stages 2-8. Y: Yolk. C4: C4 asexual adults. SX: Mature sexual adults
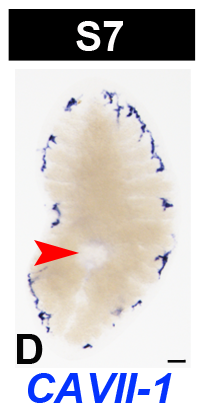
In Situ Hybridization Data
| Abbreviation or symbol | Definition |
|---|---|
| O | oral hemisphere |
| A | aboral hemisphere |
| D | dorsal |
| V | ventral |
| L | lateral |
| black arrowhead | embryonic pharynx |
| red arrowhead | definitive pharynx |
| black arrows | primitive gut |
| yellow arrows | primitive ectoderm cells |
| cyan arrows | brain |
| cyan arrowheads | nerve cords |
| blue arrowheads | eye progenitors (trail cells) |
| purple arrowheads | eyes |
| scale bar | 100 µm |
>SMED30007406 ID=SMED30007406|Name=SMEDWI-1|organism=Schmidtea mediterranea sexual|type=transcript|length=2730bp
AATCGAATACATAAAAATTTATTTGACACCAAATACAAAGAGACATTAATCTACAAGTAAAATAAGCGTTCGCGAATTCT
GTCATTTATCGCTGGAGGAGTAACACCACGATGAATCTTGCCAACGAGTTCAGCCAAACGATGGGCATAATGCGTTGGAA
CAGGAACTCGGATTGTTCCCATCCAGTTAAAATACAAATGTGTTAACGAATAAGTTATTTGCTGCAATACACTCGGAGAC
ATAACTGAAACCTCATTTGTTTTCTTGTTTGTAAACTTAGTGTCCATAAGAACATTATAATTCGTAGGAGATGCTGTACC
TTTAGTTGTTTTCTGAGACACCAAGTAAAATTCATAGAAATTGGGTTTCACTATTTTCTCATCAACCACAGTACCAGGAT
TTGGATTAGCCCCATCTTTGAAAAATTTAACACTAATTCTCTTCTTAACCACAACATAAATAATTTGGGGCAAAGTTTGA
TTTTCGTAAATTTTCTCTATCATTTTCATAACAGCATCAGTTTCGAACTTTTTGGTAAATGCTAACTGTGAATCTCCAAC
ACCGTCACGATAAATAATCAAACGAAGAGGCATAGTGTTAAATTTGTTTTGGAATGTTGTAAGGGCCGTTAGGAAGTTTT
TGCCCAAATTTTCGTGAAATTCATTTTTACCTTTAGATGAATTGACAAAACTGATATACTGAGAAAATTTGGCACTAATT
GAAAATACGGATGCTTGAACAGATCTTTTACCGGTTTTGCTATGAAAAGTATCCAATCCTACAATCATAGTCGGTGCCAT
TTTAAGATTAATACCCCATGGGTCGTATCCTAATTTAGAAAACATTTGCATAACTGATTTATCTGCAACCGTTTTACGAC
GTCGATCATCTCTATTACTTCCGTTTCTCTGCGTCACACACTGAGTCAATAGACCTGTAGACATTGTAAAATTCTTTACT
TTTGCGTAAACTTTATCATCAGGAATGAACACTAATGCCATATGTACCTTGCTTACACGAATAAAATCTGTTAAAGCGTC
TTCAACTTCATTGGATCGACATGAAGATATACTGTCCATTGTGTAATTGATTCTTAATCTTCCTATTTCTCTTTTAACAT
CTTCAATAAAATTGTTGAAGTGACTAGGGTTTCGATCAACAACCAGTACACCAAAACGATGAGCTGTGCCTTTAGGAACG
TCAAATTTAATTTCCCCAAACTTCCAATCATCAGGGATTTTAGTATTAATTTTAACTTCGTTTGAAAATATATCAACTGG
AGCCAATTCACGACCACGAATCGTAATCGGTTGTTCTTCAATACACATCCCCCAGCTCTTCATATAATCTTTTGATTTCC
CAGCTCCGGTCCACTTACAAAAATCTCTTATGTCATTTAATCGTTCTCTAGGCTCACGTTTCAATACGGTTCCTAAATTC
TTTTGCAAATTTATATTACTTCTTTCACTATCCGAAAATCCACATAAATAGCATAATTCACCTGGAATTGATAAACTTTG
ATCTGTTTCTGTAGGTCCTTCAGGTTCATTTTCAGTTTTTTTCTTTTGGGCGAACCATCGATCGCTTGACTCGCGATATA
ACAAATGGTTGTTTGTCATTTTTCAAATTAATGTTATACCGTTGCTTGAAATAATTCAAATAGGATATCTTTTTATCGCC
TAGCTGAACCTCGTCATTAACATCCATTTCTTTAATATCTGAAATTCTATACGTCTTGTTATTATACTTGGTAATTATGT
CCCTTCCTATTAATTGATCAATTTGATCTCGTCGATACACTGATACTACATTGTTTTCATTTAGAACACGATTAATTCTT
TCAAGATAAAGTAAAGTCGAAGCAGTTCCATTTTCATTTGTTCCTCTTGTAATTGTAGTAGAATACCCTCCTAATATCTT
GAAATTGTCCATTTTAAAAAAGCTAGGTCCTTCTCGAATACCACCACTTTCACAATTTGATCCCAGAAAAAATTGTCTAC
CAATTTTTTCTTGTCCCATAAATATTTGAATATTATTAAATACAATATTTATCATCTGATAATATTCTTCAGAATTTTTT
AAAAACGTCGTCAGCAATCGTATGGAAATTTCAACATCTTTGATAACTAAGTTTTCTGTATCTTCATCTGAGTGCCACTT
AGAAGTCGTAAATAATCTTCTGCCATCAAAACAAAATGGTTGACCATTATTTTTGCTGTTTGCATAAGCTAATAATAACT
CTAGCTTAGTACGGATACTTTTATTCTCCATAACTACATCATACATGTAAAATGGTCCTTCGACTTTAATATTAAAGTTA
GCGTAGTTAGACTTGAGACTAACTACTCTTCCATCTTTCCCTATTTTGTCTGTAACTGTATCAGGCTGTAAAGCAGGTTC
AATAATTGCGAATCTTCTTTGGAGACCTCCTCTTCTTGGTCTTAAATTGGGATCACTTGTAACATTAGTTGAATCCATTT
CGTAAATTTGAGTAAAAATGTTAAAACAAAGACTTCGGTTTAAATAAACAAAATATACTATGTAAAACGATAAATATAAT
TGCTTTAATGGTTTATGATTTGTATTATATAATAATAATAATTTAATTTCCCTGTGAATATTATATTAAAATATAGTAAA
AATTACTCGTAAATGTGTGTTAAAGTAATAGGATTAAATTAAAATAAACATTTTACGAAATAAGTTCTAGTCAATTATGA
AGTCTTTTCT
>SMED30004304 ID=SMED30004304|Name=Proliferating cell nuclear antigen|organism=Schmidtea mediterranea sexual|type=transcript|length=907bp
CTAAAAGGTAATAATTTAAATTTAATTAGAACATTGTTTTAATATTCGGTTATATTCTCCATTATTCTAAATTATGTTTG
AAGCAAAATTAATAAAAGGCGAAGTTTGGAAAAAGGTTATAGAAGCTATAAAAGACTTAGTACAAGAAGCAACATTGGAT
TGCTCAGAAAATGGAATTAGTCTTCAAGCCATGGATACATCTCATGTGTCCTTAGTTAGTATGCTTCTTCGAAGTGATGG
TTTTGAGACTTATCGATGTGATAGAAATGTCAATCTCGGTCTTAATATAACAAGTGTTTCAAAAATTTTAAAGGCGCTTG
GTAGTAATGATTCCCTAACAATGAAAGCCGCTGATTCAAGTGAAACAATTTCGCTTTTGATAGAATCTGCAAATGAAAAT
GAATTATCTGAATACGAAATAAAATTAATGGACTTGGATGGAGATCACTTAGGTATACCAGATACTGAATATAAGTGCAT
TGTGAAAATGCCGAGTTCAAAGTTACAAAGAATATGTCGAGATTTAAGCCAAATTGGAGAAGCAATCACGATTTGTGTTA
CTAAAGATGGTGTAACTTTTGCTGCTGCTGGAGATCTAGGAAAAGGAGCTATAAAACTTAATCAAAATTCTAGTGCTGAC
AAAGAAAATGAAGGAGTAACTATTGAAATGACCGAACCTGTTTCGATGACATATTCATTGAGGTATTTCAATATGTTTAC
AAAGGCTGCACCTCTTTCTTCTCAAGTATCTCTGTCGTTGACTGAAAATGTTCCAGCAGTTATTGAGTTCATAATCGAAG
ATTTGGGATTTATTCGATATTACCTAGCTCCTAAAATCGAAGACGACGAGTGAAAGTTTAACTATTAATGTTGTATGTTT
AATAGATGTTATTTTGTTTAAAAAAAA
>SMED30019097 ID=SMED30019097|Name=SOXP-1|organism=Schmidtea mediterranea sexual|type=transcript|length=1596bp
GGAATAACAGCCAATTAGGGATCAGCTGCAATTTGGCCTATGATTGGACATTGATTTTTATAGTCTCAGGCGTTCTCAGC
TTACGTGTGCACCTTTTTAGTGGACACGTTAAATTTGATAGGTGCTTAGTGGTGTTGATTCCTGTTTGGAAACTGTATAT
AACCAAGGTTATTTGTTGCTTTGAGACAAATGCAACACAATCAAATAATTAATTATGCTATGTTCAAATTGTTCAAACAA
TCAGGATACTTTGAAATTATGGATGGTCCATTTAGCTTCTTGTCACCCATTACCTCCTGAATGGCTTAGTGAAAAGGCCA
AAGTATTAAATGATGACGAAAATCAGCGCCCCTATACAATTGTCACTGATTCCCGAAAAAGAAGGCGACGTCCAACCACA
CCAAGACCACTAAATACGTTCATGATTTTTTCGAGGCATATCAGACGAAATATTCTCTCAGTGTTTCCGGATGCATCAAA
TTCCCAATTAAGCAAAGAGCTTGGAATTTTATGGAAGAAAGTCACAAATGAAATTAAATCCCTTTATGAAGAAGAGGCAA
AGCGATTGGCTCAATTACATGCTCTTGAATTTCCCGATTATAAATATCAACCAAAGAAACGAGTCCGAAGATTTGGAAGA
GCGATTTATGATCGACAGCCCTGTACAACAGAAAGATCTAACTCAACTAGTGCAACAGACGAAGAAAATGGTAAAGACAA
TAATAGTGTTCAAGACGGTAGAAGATCAAGCGATATTGAGAATCAGTATATAAGATATTCAGTTTTGGGTAAAAATATTG
TAAAACGAGAATGTCAGTCCTTTAATAGTCCTTTAAAAAGCGACAACGTTAAATTGGAAACGCAAGAACAAGATTTTATC
CAAGCGATGCCCAAACCTCTAAACGTCCGAATAACGAATTACAACGAAAACAGTATATTTTTACGACAACAAAGATTACC
ATCAAGTAGCACAATGTATTTACTTCCTTCAGAAAACCTAATAATTAGTAATGGGATTATGGGATCACAGGACTTGGGTG
AGGATTATTGTAGAGAGTTGTATATAATTAATGATTTTTTCCCAGAAAATGTAAACATAGACGATGAAATTGATATAACA
GGAAACATTTTTAGCAATGATTTAGATTTAGAGTGGTCTGAAATTGATGAAAATTACAAACAAGATTTACCAAATCTTAA
TTCATTATTTCAACACTAATATATTTGATTTATAAAATATTTAAATATTTTTTATTAAACAGTCTTTTACATGTACATAT
AGTATTGGAACTCTTCTCTGACTTATAAGATAACAGAGATATTGTAATAAGATTAAATTGGTTATGTGTTTCGGTTTTTG
TAAGATTGGTAATTTTCAGAAGAATTTAGACTTTTTTCATGGAAATTGGAATTGAGAAACAAACTGAATAAAGAAACAAA
TTTTTGAAAAAAACGAAACAGATACTTATTATCAAAAGAAAATTATCTGAAGCATCGAGTCGGTATGAAAAATAAACTAA
TTAGTCAGAAAATTTTTGATGAAATAACGTCTAATGAAATTAAGATCTCCAAGGAATACTTTTTCTAACATTTGAT
>SMED30026612 ID=SMED30026612|Name=Intermediate filament protein|organism=Schmidtea mediterranea sexual|type=transcript|length=2359bp
TCTTATAAATACAACATTAATTGTATAGTGAACTATGTTAGCCATTGAAATGGAGACAAGTAAAAAAATCGCAGTCAAAA
CTACGAAACAAGAAGTAAGTGGAAGTGGAAGCTCTGGTGGACTAAATGAACAACAAGCTTTTGGAAGTGGTAGCTTTCGA
ACCGGGTCTGGAGGTGTACAGATAAGTGCTAGTGGAGCAATTGGATCTAGCAGTTCAGGCGGAGTTAGTGGCACTGGCGG
AGTTGGTAGCATTGGTGGAGGCAGTGGTGCTGGTGGTGAAGGAACAGGAGCTAGCACAGTTATCACTACAAGCAGATCTT
ATTATAAGCCTATTAGTCAACCTAGAAATAATTTAAAAGACTTACAAAAAACTTCTGGTATGAGTTACAGTGCACAATCA
AACTCCCAGCAAGGTTCAAGTAATATTCCATTGATTAGGGGAGACAGTGTTGCTCTACAATCTGGAGTAACTTCAATGAA
AAAAGGACGAGCTCAAGAGAAAAGAGATCTTCAGGATCTGAATGAAAGATTCGCTAGTTACATTGAAAAAGCTCGATTTC
TAGAAGCGCAAAATAAACAATTGGTCAATGAAATTGAAAAGTTAAAAGGTGGAACCGTCCAGGACAACTCAAAGATCAAA
GAAATCTATGAAGTCGAAATCAATCAACTTCGAACTTTGTTAAGTGATTCTGAAAAAAGCAGAGCACCCTTAGAAGTACA
AGTTGATTCTCTTGAATACGAAATTGATGATTTGAGAAAACGATTAATGATGCTGGAAGACATAAACTCTCAGTACAAAT
CCACCATCGATAAGCAACGCAAAGACATTTCTGGATTTGAAGGGGAATTGGTGGAGCTCAGAAGATTCAAAGGCACCCAA
GGTGCTGATGTAAATAAAGAAAAGGAAGAAGTAAAAACACTAAGAGACGATTTGAAAAAATTGAGACAAGATTATGAGAA
AGAAGCTCTAGCTCATCTCACCCTTGAACATGAATTACAAAGCTTGAAAGAAGCATTAGAATTCGAGAAACAAATACATG
ACACAGAAGTAAAAGAGCTGGCAACCGTTGCATATCGAGACCCAACGAAAGAAAACATGGTTTACTGGCAAAGCGAAATG
GGACGCGCGTTACGAGACATTCAAAATGAATATGAAAACAAGCTTACAGGAATCAAACAAGAATTGGATTCGTCATATGG
CTCACAACTGCAGGAAATACAAACAAGCAAGACCAGGGATAATGTGGAAACGACCCATTACAAAGAAGAGAACAGCCGGG
TAAAATCTCAAATCCAAAGCATGAGATCGAGAATCGCTGATCTTGAGATGCAGAATGCTGAATACGAACACAACTTGAAT
AATATTCGTGCTAATTACGAAGAAACTCAAAGAAGTCTTGAATTGCAGTTGTTGAATTTGCAAGAGCAATTGACAGGAAC
TCAACAGGACTTGACTATTACTTTGACCGAACTCCAAAGCCTATTGGACGCCAAATTGGGACTGGAATTAGAAATCGCTG
CATACTCAAAGTTGTTAGAATTCGAAGAGAATCGTGTTAATTTTTCATCAGATGGTAACACATCACAATTTATCAGTTTT
TCATCTGGCGTGGGAGCTGGTGGTGTAGGAGGAGGATCAAGTGGAGGGTCAGGTAGAGGATCAGGTGGAGGGTCAGGTGG
TGGGTCAGGTGGTGGATCAGGTGGAGGATCAGGTGGGGGATTTGGAGGTGGAGCAGGATTAGGATCAGGCGGAGGAGGAG
GATCAGGAAGTGGGTCTGGTGGAGGCGCAGGTGGGGCTGGTTCTAGTATTGGAATTGGAGGTGGGGGCAATTTTGTAGGT
GGTTCTCTGAGATTTAACATTAATGAATTGAGCAGTACAGAGGCTGAAGAATACCAATTGAAAAGAATGAAGTTCAGACG
AACAACTAAAGGAAACATATCAATAGCAGAAGCTGACATCAATGGACTGTTTATAGTTCTAGAGAATAACGGAACAAAAT
TTGAATCTTTAACTGGAATGACGATTAAGAGAAATATTGATGGGAATGAGGATGCGGTTTCATTCACCATTCAAAACCGA
ACAAGTTTGAGTTCTGGAGAAAAACTAACTATCTGGGCCAAAGGAAAGAAAACTGGCAAAGGAAAAGAACTTGAAATAGA
AGAAGACGGCTGGGGGCTGGGGGATACTATTGTGACCTCTTTATACAATGCGGAAGGAGAAGAGAAAGCGGTCCATTATC
AAGAAATACAGGGTTGAGCAGGTGCTCACTTAAGACTGACTTGAAATTTATTTGTTTAAATAATACTTTCAATAGGGATT
TTAATGCTAATAAAAAGTATTTAAACTATTAAAAAAAAA
>SMED30014446 ID=SMED30014446|Name=SMEDWI-2|organism=Schmidtea mediterranea sexual|type=transcript|length=4773bp
ATTTAATTTTCGAAAAAAAATATGGAGGAAATACCAGTAAAAGTAGCTAAACTTGAAGCGAATGGAAGTGAAACAAGAGG
TAAAATGGGAAATCGTGGTGGCCTGAGAAGAACATTTAGAATTGTTGAACCTAACTTGCAACCTGCTGATATCACTGAAA
AAACAGGAAACATAGGAAGAACAGTTAAATTACAATCAAATTTTACCAAATTCTCTGTTTTGCGTTCAGAAAAGTTCATG
ATGTATGATTCCGTGTTTTCTCAAGTTGGTTTGTCACCCAAAGTAAAATTACAGTTTATTGTTCGAGTCTGCCAAGAAAA
TAAATTGCCAGGGGCTTTTTGTTTTGATGGAAGACGTCTATTTACTAGTGAAAAATGGCATGGAGAATCGGACATTCAAG
AATTTAGTCATGATGAAAAAAAAATGACACTAAGATTGGTATCAACTATCCTTCCTGAAACTGAAGAATATTACCAAATG
ATCAATGTATTACTCAATAATTTACAAGTAATGTTAGGTCAAGAAAGAATTGGTAAAGGTTATTTTTTAAGCCCAAATAT
TACAAAAGATGAAGTTCCCCGTAATTCTGGACCGACATTTTTTGAAACCAATAGTTTTAAAGTTCTTGGGGGCTTTGGTA
CAACTTTACAAAGAGGCACAAAACCAACTGGTGAATTAACCACTCTTTTATATATTGAGCGTATTAACAGAGTATTAAAC
GATAACTCTGTTATGAAGGCATACAACAGAAGAAGTATCGATCAGTTGATAGGCAGAGATATCATAACAAAATATAATAA
CAAAACTTATAGGATAAGTGAAATAAAAGAAATGAACGTAGATGAGAAATTTGAAATGGGTGGTAGAACTTTATCGTATG
CCGAATATTTTAAGGAACGATATAATATTCGATTAACTCAAGGCGATCAACCATTTGTTCTAACGCGAGTGAAGAAACCA
ATGCGTCGAGAAAGAAAGAAAAAAGACGAAGAAGGAGTGGAAAAAGAGAAAGAAAAAGAACGCGAAAAAGAGGCTCCAGA
AGAGAAGGACATGACTCTCAATATTCCTGGAGAGTTGTGTTTCTTATGTGGGTTTTCAGATCAAGAGAAATCGAATATGG
ATCTGCAGAAGAATTTAGGGTGCGTTTTAAAAAGAGAGCCACGTGAAAGATTGGATGATATACCAGCTTATTGTAACTGG
ATTAAAAATAGTGATGCGGCAACTGGAATGTGCAATAAATGGCAGCTGAAAATAGATAACAAACCATTAGAAATCGAAGG
TAGAGAATTACCTCCATGCGATGTAATAAGTGGTGGCAGCAAAATTAATGAAAAAATGGGAGATGATTGGAAATTTGGTC
GAGTTCAATTTGATATTAAGCGTGATCGTAAGCATGAAATCGATGTGGTCATCGTTGATCGAAATGACTTTCAGTATAAA
AATTTTATGAATGATGTGGAACAAGAATTAAGAAACATGAGAATTGATGCTCGAGTTGGAAAGGTTAATACTTGTGGTCC
AAATGATGTTGAAAGATGTTTGAATGATGCCGCTAGAAGTGGAAGTGGGTGTGCTAAAATGGCGCTGGTATTCGTCCCAG
ATGATAGAGTATATGCCAAGGTGAAATCATTTACCATGAGTACTGGTCTTTTAACCCAATGTGTAACAACACGTAATGGT
ACTAATAGAAACGATAAACGAAGAAAGGTGGTTTCAAGTAAAACCGTGATGCAAATATTTGCAAAATTCGGATATGATCC
ATGGACAGTTGAAATTAAATTGAGACCAACTATGATTGTCGGTATGGATACCTATCATAACAAGTCCAGCAAGAAATCAA
TTCAAGCATCTGTATTTTCGATCAACAGCACATTTACACAATATATGAGCTTTGTTAATTCTCCTAAAGGACGTCAGGAA
TTCCACGAAACTTTAGGCAAAAATTTCAATTTAGCGTTAGAAGATTTCAAGAAACGTTATGACATATTGCCCCAGCGAAT
TTTAGTCTTTAGAGATGGTGTTGGAGATAACCAACTTCAATTTACTAAAAATTTTGAAGTTGATGCAATGAAACCGCTTA
TTGAAAATATATACAAAGGCAACGGATTCCCTGTTCCACAAATCATTTATGTCATCGTAAAGAAAAGAGTCGGCACTAAG
CTTTTCAACAGAGGAAATAATCCAAATCCTGGAACTGTTGTGGATAAAGAAATTGTCAAACCGAATTTCTATGAATTTTT
CCTAGTTTCACAAAGAACAACGAAAGGAACTGCAACTCCAACTAATTATAATGTTTTAGAAGATACTCGATTGCTAACGA
AAAAAGGTACTATGGATCCTATGGCACCGAATGAGCTGCAAAAGATTACCTATGCTCTAACTCATTTATATTTCAATTGG
ATGGGAACTATCCGAGTTCCGGTTCCAGTGCATTATGCGCATAGATTGGCTGAATTAGTTGGAAAGGTGCATAAGGGCTC
TAATCCCGTTGCCATTAATGAAAGAATAAGAAATAGGCTCTTCTATTTATAAAAAGATCAAAACTTACATTTTTCTGTGT
TGATTACTATCCCAATTGTGGATTTAATTGTTTCTGAAAGCTGTTCATTCAAATTCTTCTGAAATCCATATTGATTCGTA
AGCAGCCTGTATTTGAATACTAAAATTGATAAAGTAGTTATAAGTATTGTAAAAGTATTAAATATTTGTTTATGTTATTC
GTAATACACTTGGGTTATTGCAATGTTTTAGTTGTTTTTCAATGATAATAAAGCTTAGCTGTATAAATTCTTTTAGGTTT
TTTAGAAGGTTTATTTATATCGTGTAATTGATTGTCGTGTTAATATTAATTAAATAAAAGTTTGATGTAATTGCTTTGTG
TTACGCATAATAAATCTTACGAAATTTCTGAATACAACTGCTGGTGATCGGGATATTGAATGCCGGGTCGTTTGAATTAA
TAATACAAATGCTTTCGATTATTGTCATTAATATTTTCTCCAGTGCGTAATGGGCCAGAGTTCCGGTGCCATGATTAATG
GTTCCTAAATTTTAGAAATAACATTTTGAATGACATGTTAAATGTGTATATAATAAATTTTCTAGTTTTTTTGCTAAGAT
TTTGAGAAATTTTAGTTACCTGTTTTTGTTAATTGCTTTGATAATTTCAATGCTGACACGTGCAGAATTTCACACATCTC
CAGCAGTCGACTGAATTCTTGGATTTTGTTTGATACAATTATGCTTGCAGATGTTATGGCGCATAGATCCAAGAAGGCTG
TGAATCTTGGGTCTTGTTTTGTAAAGGAAGCTTTGTAAAATATGATGGGACAGTTGCACATAAATTCTAGGAATCCAGAT
TTTAGGCAAGATTGAAGTAATGTTGAATGGTTCTATTATAGAATAATCAGTTGCACTTTTAGATAACCAATATACCACAA
AGAGCCATCCGCAAATTTTGAAAGCCCAAAGCATTGAATCGTTGTCTATGTTACTCAGTATAGACAGCAGTAAAATGGCG
CTTTTTTGATATTGTAAACCGAAATGATTACTAGAATTCATTGTTTCGTCAAGAGATTTCGCGAAACTCTGAATATTATC
TGATTGATTTGATACTATTCCTTGGAGTTTACAAATTGATTTCAGTTCCAAATTCATACTATTGTGTTCTGCTGATTTGC
TACTATTGGAGCTCTATAAGGAATTATTAGAATCAATTTTGGTTTGATAGTGGGTTTTGCAATCATTGGCTTATTGTGGG
TTGTGTAACCATGGAAGTAATTGTGAATAGTTATAGTGCTCGCTCTGCATAAAATAATATTTAATCAGTGCTTTAAAGTA
GATTTTTAACGATTATTATGAAAGATTTCTCTTATATATTAATGTTTTTGTATCCTTCAAAAGTTTACTAAAAATAATGC
ACATTTGAAATTTACGCGTTACCATAAAATGTCTGAAGTGGTCCAAAGCAATAAAAGCTTAAAAAGAAAGTTCTTGTTTT
TAAATGTATTGGGATATACTCCCACTAACCTTCCGTACTCCGCTGCCTCCCTTCCCATCACATACAATATTTGCTAGTTC
TACTTACAGAGTTGAAAATCGAAAAATTTTCCCAGTCAATGTCCATGATTATGCTGAATAAATTTTGTCCGAACATAATA
GAGATTGCCTCAGCCAGTATTTCTTTCGTTGTTGAGTTTTGATGAATGACATGTGAGACTAAATTTATGGACAACCATTG
ATTTTGGTTTTGCTGAAGTTGAGACTTCTTACGGTAAAGATAAACACGAAATAATGATAAACTCTAATCAAAAGATATTA
CAAACAAATATTATGATATCAAAACCTTCTGTCGAACCAAGGCACTTGTGTGAGATAATAAATGAAGAATGAATTCGAAA
AAGAAGTCATAATTCGAACATCGCTCCAGAATGCTGTCACTGACTTTGACTTCAATCACATTTTCCAATATTGAAATCAG
ACCGGCGATCACATGGACCCTTGATGATTCAGACTGACCACTGATTACGGGTTGATCGGTTAAATTATCTAGAGATTAAA
ATAATAGTCGGTAAAGGAAATTATCCATATAAACGGGAAACAGAAAAGTTTTAACCATGGTTTCAAAAGCTTTTGGGACT
ACCCCCAGCTTTGCAAAACAATTTTTTAACGCTCTTGTTATGACTTGAATCAACATTGTACAATTGTTACAATAGACTTG
ATGATTTACTTATTTACACCCCATCTAAATTGTTTCTTAACCCTTTGCCACCC
>SMED30018353 ID=SMED30018353|Name=Glycine amidinotransferase, mitochondrial|organism=Schmidtea mediterranea sexual|type=transcript|length=1408bp
CAAGGGTAGTTTGATTATTTAGAAAAAAAGGATTTCCACCGGTTTTCTGTGATAAAAATTGTACAAAAATGCTTACCAAA
ATTGCTAGATCTCTGCACGATCAGATTTCCATAAACGTTGTCAACAAATTGTCAGCTAGGTTATGCTCGTATACACCTGG
TAACAAAAAACACAGCCCGGTTTGGGCCTGGAATGAATGGGACCCATTAGAAGAAATCGTTTTGGGACTTCCAGATATGG
CATGTTGTCCAGAGATTTTACCAGAAGCCAAAGCTTGCATTTCGGAAAACAAATTTGATTTCCTTTTAAAACACAAAGGA
AAATATTGGAAGGATGTAATTGAACCAGAACACTACAAAATAATGCTAGATGAGCACAAGAACTTCATAAAGATTCTACA
AGGAGAAGGAGTTAAAGTCGTCCATCCAGAACCGATTGATTTCGCTAAAACGCTCTGTACTCCAAACTTTACTGCCCATG
GAGTCGACTGTGCGATGCCGAGAGATTTTCTATTAATTGTCGGAAATGAAATGATTGAGTCCACCATGACTTGGAGATCG
AGATATTTTGAATTTATGTGCTATAAAAAATTAATTAAAGATTACTGGAGCCGGGGCGCTAAATGGACAGCGGCACCAAA
GCCGATGTGTAAAGAAACACTTTTTAAAGAAAATTATCGTGCTGACGAAGATTCAACAGCATTTTTCGATGAAAAGACTT
CGATTCTAACGGAAGAGGAACCAGTTTTCGACGCCGCAGATTTCATTCGATGTGGTAGGGATATATTTGCGCATGTCAGT
CAGGTTAGCAACAATGCAGGAATTGAATGGGTACGTAGACATTTGGCACCGAAAGGAATTAGAGTTCATTCAATGAATTT
CGTTGACCCTAAACCAATGCATATTGACACGACACTTTTCACACCTCGGCCAGGCCTCGCGATAACATGCCCAGAAAGAC
CATGCTTACAAATTGGTATGCTTAAAAAAGCCGGCTGGGATGTTGTCGTTGCTCCTTATCCAGATATTGCAAAGAATTTC
CCATACTATATCAGCTCTCAATGGTTGTCTCTTAATGTTGTCATGTTGGATGAAAACCGAGTTCTCGTAGAGAAAGATGA
AATTGGAATTCAAAAACTGTTTGAAAAGTGTGGCATTACTCCTGTAAAAGTTCCATTCAAACACTGCTTTTCCATAGGAG
GTGGTATTCATTGCTGGACATCGGATGTTAGAAGGCGAGGAGACCTTCAAAACTATTTTGACTGGCCTCAAGAAATCTTG
GAGAAATTTGAAACTCACTGAATATCACAATTGAAAATATAAAGAACATAACTTTTCCAACGTATATTAATTATTTGTAA
TAAAATACACTGCTTACATCGTGTCAATTTTATTTACTGATTAATTAC
>SMED30001900 ID=SMED30001900|Name=Bruno-like protein|organism=Schmidtea mediterranea sexual|type=transcript|length=1713bp
CTTTTGAGGGAAAAATAATTCTTTTATTAATAATGTTTCATCATTGGAATATAAAATAAAATATTAGGAGAAAACATAAG
TTAACCAACAATATTAGCCACATATTTCGTTTAATTATTGGTCTATCACTTTTCATTTTAACCAATTGTCTTGAGATAAT
GGAGAGTTTATCCTTAATTTTTATTATAAATCACTGGTTTTGATAGACTTATTAACAATGGAAAATTATTTCAAACTTTT
CGTTGGTCAGATACCTCGAAATACACCAGATAAAGAAATTAGACTTATTTTTGAAGCTTTTGGACCAATCTTTGAATTTT
TAATTTTAAGGGACAAGAAAACTGGTTTTCATAAAGGATGTGCTTTTGTTACATTTTATCATAAAGACTCTGCAGAAAAT
TGTCAAAAGGCGTTACATGACAAGAAAATCCTAAACGGGATGAATCGACCACTTCAAGTGAAACCAGCTAAAAATAAGTT
AGCAAAAAATTCAAGTTTGAATGGGGAAAATCAGACAGAAGAGCAGGTTTTCAGTGATGAAAGGAAATTGTTTGTTGGAA
TGTTGAGTAAAAATCAAACTGATGAAAATGTACAAAATATGTTCACAAAATTTGGCAAAATCGAAGAGTGCACTGTATTG
AAAGACCAAAATGGAAATAGTAAAGGATGTGCCTTTGTCAAATTCGTGAACCACACCGATGCTTGTGCAGCAATAAACGC
TCTTCATGCCAGTCAAAAAATGGAGGGTGCTTCATCCAGCCTTGTGGTTAAATTTGCTGACACGGATAAACAAAAACAAA
TAAGAAAATTGCAGCAATATATTCCAGATTTAAGCATTTTAAATTCGCGCATTCCTATTAATATTCCATACTATTCACAA
TACGGTACAATAATTAATCCAGAATATAATTCAACAATTCCTTCAAACCAATATATTCGAAATTGTAACGAATATATTCT
TGCTGCTCAAATTCAACAAATAAATTTGTTTTCTAATTTATCATCGGTCTACCCATCTAATTCAATAGTGTCAATGCCAT
CCAGCCATCCGGTACCAATGATTGACTATTCTGTTCCTATTAACAACTTTCCACAAACTGACCAAATAACAATGAAACCA
ATTGTTGATATACAGCCAATACCTCCAGAAGCTGTACACCGAACCATCGTAACTCCAGGATCACTCTTATCATATCCTTT
TCTTAATGTTCAAATTCCTACAGTATATCCAAACGCCTTTTCCAATGGAAATTTAACAAGTACAACACAAAGTTCAACAG
ATGTGAAAACCACTCCTTTGCCAACAGGTCCTGAAGGCTGCAATCTATTTATTTATCATCTGCCACAAGAAATTGGTGAT
TTACAATTATATCAAATTTTTATGCATTTTGGAAACGTTATCAGTTCTAAAGTTTATGTTGACAGAGCAACAAATCAAAG
CAAATGTTTTGGATTTGTCAGCTACGATGATCCGGCATGTGCCAATGCCGCCATCAAATCTATGAATGGATATCATATCG
GAACAAAAAGATTAAAAGTTCAGTTAAAAAAACCAAAAGAAGCAATAAACGAAATGAAAAACTTCAATCAAATGTTATCA
TCAACTCCCGACATAAACCAAGTATTTTGTTCATCCTAATGATTGTGATATTTTACAGGATTTTCTCGACCATATTGAAA
GCCTGATTGGTGATATGTAAAATGACCCACCGT
>SMED30008552 ID=SMED30008552|Name=Carbonic anhydrase 7|organism=Schmidtea mediterranea sexual|type=transcript|length=911bp
CCGATCTAAATAATTTAATTCAATAAATAATTTCAACTAAAGTTTTTATGATTTCTGAAGATTGGGGATATGGACCTGAG
AACGGTCCACATTTATGGCACAAATCGTATTCCAGTGCGGGAGGATGTTCCCAATCTCCTATCAATATAGATCTTTCACA
GACAAAATACGATATGACTTTAAATTTACATCCTTTAAAAATATGCATTGATAGTTCGAGGACATCAAGTCAGACTTTGG
TATTTACAGGACATAATTTCCGAATCGACTGTGAAGGAGCACTATGCGCATCGGGAGGCCCAATTCCCTATGGTCAAATG
TACAAAATTTCCCAAATTGTGTTTCATTGGGGCAGTACAATTTATGGAGGATCTGAACATTTGATAAATTGTTTTCAATT
CCCATGTGAATGCCAAATGACCTGCTACAATGCCACTCGTTATAGGTGCTTCAATGAGGCGGTAAATAAAAGAGATGGGC
TAATTATCCTATCTACAATGATATTGGGAACAAAAATAAAACCCTCGATCGTTTATTGGGTCCTTTGACGTCGATTCACT
TTTTCAAACACCATGACTTCCGGCAAGAAACCGAACTGTTAGATTTGAAATGTCTTTTACCAGATTGCATCAATCGATAC
TTCACCTACACTGGGTCATTGACTACCCCACCTTGCTATGAAACTGTTACGTGGATCATATTCGAAGGATCCAACACAAT
CAGTGAGGATCAACTAAATCTCATGCAGAAAGTTCACGTAGCCGACGGCTGTAATGGCTTTGACAACTTCAGACCGGTTT
GCTCACTCGGCCAAAGAATCATTACCAGAAATTTTGTTTGAGCCTTTTTTGAACTTGATTTATTTAAATTTTATTTTTTT
AATTGCTACTGTAGTGGATGTTATGGGGAAT
>SMED30020435 ID=SMED30020435|Name=Myosin heavy chain|organism=Schmidtea mediterranea sexual|type=transcript|length=3615bp
TCTGAATTTAAACAAAGATATTCAATTTTGGCCGCAGATGTAATAGGGCAAGGATTTATTGATGGAAAACAGTGTACTGC
AAAGATAATGGAAGCAATCCAATTAGATAAAAACAACTACAGGCTTGGATCAACTAAAATCTTTTTCAAGGCCGGAGTAC
TTGCTCAGTTAGAAGACATGCGAGATGAGAAATTATCATCCATTTTGTCTGTATTCCAAGCACAGATGAGAGGATATATA
ATGAGAAAAGCATATAAGACATTGCAAGATCAACGAATTGCTCTTTCTCTCATTCAAAGAAACATCAGAAAATATTTATT
ACTAAAAACATGGCCATGGTGGAGACTATACACTAAAGTCAAACCACTCCTCAATATCGCAAGACAAGAAGAGGAGATGA
AGAAAGCTGCTGAAGAGTTGATTAAATTGAAAGAAGAATATGAAAAGATAGAAAAATTTAAAAAGGAATTGGAAGGTCAA
AATGTGGTTTTGCTTGAGCAGAAAAATGATTTATTTTTACAACTGCAAACAGAACAAGACAATCTTAGTGACGCTGAAGA
AAAGATTCAAAAGATGATCAATCAAAGAGTTGAAATGGAGAACCAATTTAAAGAATTAGAAGATAAATTAATGGATTCGG
AAGACATTCAAGATGAATTGGAAATCGAAAAAAAGAAATTGTTGTTGGATATGGATGAATTGAAATCGGACATTGACGAA
CTTGAAGTATCTTTAGAAAAGTCTGAACAAGAAAAACAAGCTCGCGATCAACAAATTAATAATTTGAAAGAAGAAATTCA
AAAACAAGATACTTCTATCGGAAAAGTTCAAAAGGAGAAAAAGACTCTAGAAGATAACTTAAATCAGACCAGAGAAGCTT
TAAATGCTGAAGAAGACAAGGTCAATCAGTTGACTAAAATCAAATCAAAATTGGAATCTGAAATAGATGAAATGGAAGAA
GCTCTAGAACGGGAAAAGAAGATCCGTGGAGATGTTGAAAAACTTAAACGGAAATTAGAAGCTGATCTCAAATCAAGCCT
TGATAACTGTAATGAGTTGGAAAATATAAAGCAGGAGTTAGAAGAAAATATTCAGAGAAAGGACATTGAAATATCAAAAT
TAAATGTCAAATTGGAAGATGAATCTTCAATTGTCACTCAACTTCAACGAAAAATTAAGGAATTACAAACTGTTATACAA
GATCTTGAAGAAGATCTAGAAGGTGAGAGACAAGGTCGATCAAAATCTGAAAAAGCTCGACAGTTGGCTGAACAAGAACT
CGATGAACTAGCTGACAGACTTGAAGAGCAGGGAGGTGCTACTATGGCACAAGTGGAATTGGCCAAGAAACGAGAAACTG
AATTAATGAAACTTAAAAAAGAATTACAAGATACCAATTTACAGAATGAGCAAATACTCAATGGTTTGAAAAAGAAAAAC
CAAGAAATTACCAATGAATTCTCCGAGCAAGTCGATCAAATAAATAAATCTAAATCAAAAATCGAAAAAGAAAAGGCGAC
TCTTAAGCATGATCTGGATGAATTTGTTTCTCAATTAGAATCTGTTTCAAAGCTAAAGGCCAATACAGAAAAACATTCAA
AACATTTAGAAAGTCAAATTATGGAATTGGAAAACAAAATAGACGAACAGAATCATCAAATATCAGAAATGGTTTCACAG
TCTAATAAAGACTCTCAAGCAATAATGTCTTTAGAACAACAACTAGAACAAGCAGAAAATGAAAATCACCAATTGAGCAA
ATCCAAACAACAATTCCTCCAACAAATTAATGACTGTAAATCGACTCTTGAAGACGAAGAAAAAGTAAAGGCCAAACTTT
TCAATGAAATTCGTAATCTGAATGATGAAATGGGCAATTTAAGATCCAGTTTGGAAGAAGAGCAAGAATCCAAAAGTGAA
ATTCAAAGGTCACTAGCAAAATCCCAATCTGAACTTGTACAAATGAAAAATAAATTTGATAGTGAAGGTTCAGGTCGCAT
TGAAGAAGTTGAAGAAGCTAAACGTAAGTTGCAAACGAAACTTCAAGACATGGAACAAGAATTTGACGCTATTAAAAATA
AATGCGCGCAACTTGAAAAAATCAAAATTCGTCTCACTGGAGAAATCGAAGACATGAGTGTTGATGTTGACAGAGCTAAT
GCAGTGGCTAATCAACTAGAAAAGAAACAAAAGAACTTTGACAAAACCGTCAGTGAATGGCAGTCTAAATGCAACTGCCT
TCAAACTGAAGTTGATGTTGCAAATCAAGAATCAAGATTTTTGTCAACCGAAGTATTTAAATGTAAAAGCCAAAACGAAG
AATTGCTTTCACAAATCGAAAATCTTAAACGAGAAAATAAGTCTCTCAGTGATGAAATTCACGATTTAACGGAACAACTA
AGTGATGGCGGAAAATCTGTTCACGATATAGAAAAGGCAAAACGAAAATTGGAAATAGAAAAGGAAGAACTGGAACAAGC
TTTGGAAGAAGCTGAATCAACATTGGAAAATGAAGAAGCCAAATTACAAAGAGCTCAGTTAGAGTTGAGTCAGATGAAAT
CAGATGTAGAAAAACGATTGTGTGAAAAGGAGGAAGATTTTGAAGGAACTCGCCGTAATCATCAAAGAGCAATGGAATCA
ATGCAAGCTTCTTTGGAGGCAGAAATTAAAATTAAACAGGACTCAATTAAAGCCAAGAAGAAGTTAGAACAAGAAATCAG
TGAACTCGAAGCCAATTTAGATCAAATCCAAAGACTGAAATCAGATGCTGATAAGAACAATTCAAGAATGCAACAGCAAA
TTCATGATTTACATTCAGAGATTGAACAAGAACAACACAGTAAAAACGAAGTCCGAGAACATGCTCAGATGGCTGAAAAG
AGATTTTTATCGGCTAATGGAGAACTAGAAGATTTGCGAGCCAAATTTGAACAATCAGATAGAAATCGTAGAAATGCTGA
ATCTGAGCTTCAAGATTTGAGTGATAAATACAATGAAAGCCAAGCACTGATCACTTCACTTCAAAGTTCAAAAAGAAAAA
TGGAAATGGATTACGTAGCTATGCAGTCCGATCTTGATGATATGTCAAACCATGTTCAACAAGCTGAGGAAAGAGCAAAG
AAAGCAGCTGGTGACGCTGCCCATATGTCAGAACAACTTCATCAAGAACAAGACAATAGTTTGCACACGGATAAAATGAG
GAAGCAAATGGAAATTCAAATCAAAGAAATGCAGTGTAAACTAGAAGAAGCCGAAAGTAATGTTTTGTCAGGCGGCAAAA
AGACTTTGGCAAAGTTAGAGTCAAAAATACGAGAATTGGAAACTAATCTTGATAACGAAATGAGAAAACATGCTGAAACT
CAAAAGAATTATCGCAAAGTAGATAGACGAATGAAAGAACTAACATTACAATTGGAAGACGATAGAGCTCAACAAGAAAA
ATCAAATCAAGTTATTGAAAAATTCCAAGGACAAATCAAAACCCTTCGAAAACAATTAGAAGAATCAGAAGAAGTTGCCG
TTTTGAATATTGCCAAATTTAAGAAAGTACAATGTGATTTTGAAGAAGCCGAAGAAAGAGCGAATGAATCGGAACAAGCT
TTACAAAAATTAAGA
>SMED30024586 ID=SMED30024586|Name=Pax6A|organism=Schmidtea mediterranea sexual|type=transcript|length=1992bp
GTTGGTTTTTCTAACAGATTTATTAATGCTAGATTGGCCGCTCAAGATTAATCATAATATTCTGCTACTTTACTGATATA
ATTTTTATACACGGTTACTACTACTCCTCTACTATTTTCGCATTGTTGGCTTCTCAAATTCAATCCCTGGGCATAAATCA
AACCGCTTCATTGACAAAATCCTTTTTGACACTACAAGAAATTGATTCAGTTACTGAAGTAGAAAATATACACAGTCGAA
ACCTATGACCGAATCGAATGAAGAGGAAATAAAAAATAAAAAGAAAGTTAAACGAGGACATTCAGGAATCAATCAATTAG
GTGGAATGTTTGTCAACGGTCGACCTTTGCCCGATTCGACCCGACAGAGAATCGTTGAGTTGGCTCACTCCGGAGCGAGA
CCATGTGATATTTCCCGAATCCTCCAAGTTTCCAATGGATGCGTCAGTAAAATCCTATGCAGATACTACGAAACCGGTTC
AATTCGACCGAAAGCCATCGGAGGCAGCAAACCAAGAGTAGCTACCAGTTCAGTTGTGTCGAAAATTGCCGCCTACAAAC
GTGAATGCCCGTCGATATTTTCTTGGGAAATTCGCGATAGATTATTACAGGAAGGGGTTTGCAATCAGGATAACATTCCC
AGTGTCTCTTCGATAAATCGTGTTCTACGCAGTTTGTCTAATGAAAACCAACGACATTTAGTTGCAGCAACTGGAATGTA
TGACAAACTGTCATTGTTAAGCGGCCAGCCTTGGTCTACTGCAGCTGCTCATGCGGCTTGGTATTCCAGTGCTGCAGCAG
CACATGGGTATGCTTCATCGACATTTCCCAATTGTGGTGCCTATGGAGGCTTAACCGGAATCGGAATTATCAATGGAATG
AGTACAGCCCACGCGGTAGCGTCAATTAATCAGTCGAATTCCGGGGTCACAAATTACCATGTTCAATCAACAACCGATAG
TTCCGACAAACACAAATCAGAAAAATATTCAGAATCCATTGCACATTCTGAATCGAATGCTTCCAGTGAGCCTGGCAACG
AATATATGTCCGGAGTAAAAAGTGAGAATGATGATATGAGAATTAAACTGAAAAGGAAACTTCAAAGAAATCGAACATCG
TTTTCAACAGATCAGCTTGATTCTTTGGAAAAAGAATTTGAACGAACTCATTATCCTGATGTATTCGCCAGAGAGAAATT
GGCAGACAAAATCAGTCTTCCAGAGGCGAGAATACAAGTTTGGTTTTCAAATCGAAGAGCAAAATGGAGACGAGAAGAAA
AACTTCGTCGACAACGGCAAAATCTTATGCTTGGTTCAAACGGAACAAGTTCTACTGCTGATACTAATGTAACAACAAAT
GGAAATACTCAATGTTTGTCGACAACCGGACAAAATTCAATGGGATTTTCTGGAATAAATGATATCAGAAACCAATTTGG
TGATGTTGGAATGCAAACACCTTCATTGTCGGCTGTAGCAGCTGCAGGAATGTACCATTCGGCAGTTGCCTCTGCTGCAG
ATCAGTATGCAAAATCAGCCGGTTTACCAAATCTTTCGCAATCGTCAATCAATTCTCCATATTCCTACATGGCAAATTTG
CAGTCGAGATCCAGTGGTTCCCAAATTGACTTGCCAAACGTTACAAGCAATTCAAATTCTTTTTACAACCCCGGATCAGT
TTATCCCCCAAGTTTATTTCAAGGTGTAATGAGAGGATATGACACCATTGGATATCCCAAGTATCCTTCAGCAATGTTTC
CAGGATCACTCGAAAGTAATAATAATATCAACCCTGCCGGTTTTCTTGCAAATAATATGTCTTCGAATTCATCTAAACCC
GATTACGATTTCTCAGCATTTAATCGGCCCAGCAATTATTGGACTCCGGTCGCATAAAATTGCACAAATGAATGAAAACA
ATTTCAAAAATTCCGAAACAACAATGATTAGAACAAAAAAAACATTTTTCTAAAATAGTTGTTAATTTAAAC
>SMED30021108 ID=SMED30021108|Name=POU domain protein|organism=Schmidtea mediterranea sexual|type=transcript|length=1667bp
TTCTGGATTACGAATAGCCTATTGCAATTAATAACCGAGTTTTTTTCAAGATTGTCCTACAATGATCATTAAATAAAAAT
AATCCACATGAGTATATCAGAAAATAATAAAGAAAAAGATATTATTTACGGCTTGGAAGCAAAGCATATTGTAGATTTCA
ATGTTGCATCTCATTTAGATGAAGAACAAGACGCAATTTCAAAATCTATTGGGCGCAACCAAAAAACTGATAAACACTCT
CAGTCGTTAAATGGGAAAACTCCCAATGAACCACTGGATCACAATTCATATATCAAAGCCTATAACGATGCGCTGTTTGA
ACAGCGGACTAATTTCAATGGTTTGAATTCAGATTTAAGCTATTCATTTTTATCTCAGTATGGCTATAATAATGATGAAA
GAATGGATTTTACACCGTATTCTTTGCCAATTGATATGATTGAACAATCTTATAATTCCTGGTCTAAATCTAACGCATGC
ACATTTACAGAGGCGCATCTAAATTCACCTTGGTACAATCATTATCCGTATCCAGGATATGCATCATATAATTCTTATAT
GTGTAATATTCCACCGGTTGCAACTTCATCTATCAATGATCAATCTAATATGAGTTTTCCTAATCATCCAGCATTCTCAG
ATCTGGAAGAGTATGAACAATTAAAACAATTCCAAGCAAGTAATTATTCTGTATTTCCACAAGATTCTTTTTTAAATCAT
TCAATATCAACTCTAATGCCTCCGTTAAGGAATTCTCAAAGCACCACCCCTTTCAACTCAATCAATCCATCATCTCATTG
GTTGGATGTTAAAAATTGGCCGAAATCTGCCTCTCAATCAGATCAATTCGATCAAAATTCATTGAAACGTCAAATTGATC
ATTTTCCTTTGAATATCCCAACGGCGTTTTCTTATCAATCATCCAACTCAATGTCCCTACTCGGCTCTCAACAGCCAGAA
ATCAGTAAAGATCTCTTTTTGTCCGATTCCATAATTGAAAATTCAAAACTAAGCACGATGGACTGTCGAGAATTGGAACA
ATTTGCCAAGACTTTCAAACAAAGACGAATCAAGCTAGGATTTACTCAGGCTGATGTCGGTCTGGCGTTGGGTACGTTAT
ACGGGAACGTATTCAGTCAAACCACAATTTGCCGATTTGAAGCATTACAGTTAAGCTTTAAGAACATGAACAAGTTGCGA
CCGTTATTAAGCAAATGGTTAGAAGAGGCTGATTTGTCCTTGTCAAACACAGTCGTGAATGTAGAGAAATTAGCGAACCA
AAATCGACAAAGAAAGAAAAGAACAAATATAGAAATGAATACCAAAGATATATTGGAATCAAGTTTCAATAAAGATCCAA
AACCAACTGCTGCTCATATTATATCATTGTCAGAGAGAATTTTAATGGATAAAGAAGTAGTTCGGGTTTGGTTTTGTAAT
AGAAGACAGAAAGAAAAAAAATCCAGTTTAACCAAATCTATTTCCAGTAGTTAATACAATCTTTCCAAAAAGAAATTCTT
TTACAGATTCGACGTTTTATTATGGAAATGGTGTGAATAAATAATTTCTTCCCATAAATTTTGCCGGTTTTCTAGCTTGT
ACAGATAAAGAAAAATACCCGTTTTTACATTTTCCTAAACTTAAAATATTCAACATTAATTTTGAAC
>SMED30035063 ID=SMED30035063|Name=SOXP-2|organism=Schmidtea mediterranea sexual|type=transcript|length=2664bp
TGTACATTGAATTCAATATTTAATAGTGATTAACTTATCCAAATTTTCAGCCACTTTTAGTTTAATTTGCGTGTTTTGTT
GAGCAGTTTTTATTGGCGCATTTTATAAAAAATACAAAACCAATCCAAATCAGTTTGAATCATTGTCAGATTTTAGTCGT
GCAAGATACATTATTTCATTTTAATTCTCTAGCATATGACATTTAAACATGGAGCTGAATTACTCGTTGAATGATAATAA
TTTCAATTTACAATTAGAAAGCGAGTCCTTGACTTATTTCGATTTATGCCAAAAATGTTCAGCGTACCAAAATAGTCTTA
GAGACTGGATGAAACATTTAGTTGACAGACATCCTTTACCTATTCAGTGGATGAGTGATTTGGCAAATAATTTAACAGAT
GATAAATTATCTAAGCCATATTCGACCGTTACAGAATCCCGTAAAAAACGCAAGAAGCATATCACTCCAAGGCCATTGAA
TAGTTTCATGATATTTTCCAGACATTTAAGAAGAAATGTTCTTATTTTCTTTAGTGATGCTTCAAATTCAGAAATAAGCA
AACTTCTTGGGAAGTTATGGAATAACATTCCAGCTACCATTAAATCGATGTATGATAATGAAGCTAAAAAGCTTGCAATT
TTACACTCCAAAGAATTTCCTGAATATAAATATCAACCTAAAAAGAAAAAAAAACAATATCTAACGAAATGCCCCATTAA
TTCCACACCTTTGTCGAGTAATTCATATGAAAGTATTCAAGATATTAATTCAGTAACGGATACACTACAGAATTCATCTA
ACAACTTACAAAAGGAAAGTAATCCGACCTCTGTTTCGTCTTATGGAAGCAATCAGGAATGTAATAATTCATTTAAGAAT
CATAGTGTACAGTCCAGCAATTTTCCCAAAGAAATCAACCATAAACCAAGTAGAAAAATTGAAAAGCAATCGTGCAAAAT
AAGTTCCAATCAAATAATGGATGATTCAGAAGGGGCTGAATCCAGTCCATCATCATTTAAAAACACCATTAAAAATGAAA
TTTATTCCTCTTATAGTCCATTTCTAAATGAATCCCATAGCATTCTTTCAGATTTAAGTTACTCTTCGTCTCTCAATAAC
GTTTCAAAACACACTAGCGATTCCATTATAACGAATTCTCCTATCAGAGAAGAAATTGTCTTAGAAAATAATCAGCCCTC
TCAACAGAATAATATAAACAATTACCGACACGTATTTCAGAGAACTTCGACTCTATCCAACAGTTTGTTGACCCCTTTAA
CCATTGTCACAGAAAAAGTTGACGAACAGAAAGATCTTCCACCATCAAGAATCAATATCCGATATGTGAAAATTAACAAA
CTACCGATTCAAACAAAATCCGATTTCCCATTGACTGGATATTTTCAAAATTTCAATAACGACTCAAATCTTCATTTTCA
GAGTTCCATTCACGCCCTGAATACGACATTGACTCCAGATCTGTCGGTAAAACGCGAAGCCACTGTCAACAACAGCATGA
CAGATTATATCAATCGAAAACGTACAAATGTCGATCCACTTATCATCGATGATATATTTGAAAACGAACAATTTGAAGAT
GAGGAAGTGTTGAATTTTGACAATCCTGATAACAATAACAGTTCTCCTTGGTCGTTTAAATCGCCTAACTTTAAAATATC
TACCGGTTCCTCGACACTGTCAGACAATCCGTACAAAGAACAGTTTTGTTCTTCCCCGTTCCATCTATCAGAAACTGATA
CTCAACCGATTCCAGTTCAACGATTGCCAGCAATCGGCTCATGGTTAAAGTCAAACTATTCTGAATTCATGATCTGATCC
AATCTATTATTCTATTCAATTGTCTGTTTTCATACAATTAGGATTACTTGTTATGAATTCTAGCTGATTTTGAGTGTGTT
TATAAAGCTTTTGTAACGCCTTTTCTATACAGATGTTCGATATATATATATAAATAAATAAATATAAATATATAACAATT
TTATTGACATTGTACAGTATTCCATAAGCCCGATCTTGTAACGCATTTATTACTATACTCAATAGACCAATCTATATGTT
CTCATAATTTCTATGTTGTTATTGTACACGGGAATTATTACTATAAATGATGCGAATTTATCACTTTGTAATTGAGGGAA
ATTATCGATCAATTTGCGCTTTCAGACATTTTTCATGTATAAACTTGTTTTTCTCTTTTTTATCATCAAAAATTTAATTT
TTTTGATATCGCTATTCATCATATTATTATTCTCCGTTGCATTTGTTAGATTATGTAAATTTTTATATGAAAATGTTGTT
ATTTATTGTATTTTTATACGATTTTTTGCTTTGGATACACTTTTATCAAATCATATTGCCAGTGTGAGCATTTATTGGAC
TTGTTGTTATTGCTGCTGCTCGAATAATTGCTTCATACGGGGAGTCTCGATATTTTCATGAAGCGATTTTCATGGTTGCC
GCTCTCCGCCGCCGCCATTCGGTGTAAATATCAATTGCATTGCCAGCACGTTTTCTCAAACTCAATTTTTGTACTCAATT
TCAATTGATTTATGTTCATTTTATTTTTCATAGAAATGAGTAAATAAATTTTCATAAATAGTTTTGTTTTCAATATTTTT
AGGAAATTTGAATGTGTCTCTAGA
>SMED30022907 ID=SMED30022907|Name=Synaptotagmin|organism=Schmidtea mediterranea sexual|type=transcript|length=1814bp
ATTTATTGAAAATTATTTTTTCTATTATTAGGAGAAAATTGATTATTTTTTTTTGACAAAATAAGTTTTGGATATATTTC
CCATTTGTTTGTATTCGTAAACCAATGAAGAATTCGACTCACGGATCGGATCATTCGGCCAGCAAGGCCAGTATGAAAAC
GAATGTGGAAAAATTTTGGCATGATCTCGGAAAAAAATTGCACATGGATCCAAAAGCATTAATAGGAGTAACAATCGGTT
TGGGAATATTGTTTTTGTTGTTCCTTTATTGCTTATGTAAAAAATGTGTTTTCAAGCGTCGAAAAAAGAAGGAAACGGCC
AAAAAAGGAGCCAAGGGAGTAGTCGATATGAAAAGTGCCCAATTGTTGGGGAATTCATATAAAGAAAAAATTCAACCCGA
TCTTGAAGAACTTGATGCAAACATGGAAGACAATGAAGGTGTAAAAGAAAAAGAAGATGTTCATTTAGGAAAATTACAAT
ATTCTCTAGATTATGATTTTCAGAAAGGAGAATTAACTGTTGGAGTTATTCAAGCAACTGATCTTCCAGCTATGGATATG
TCAGGAACATCAGATCCATATGTCAAATTATACCTTCTCCCAGACAAGAAAAAGAAATTCGAAACAAAGGTTCATAGGAA
AATTTTAAATCCTGTATTCAATGAAACTTTTGTTTTCAAAGTTCCTTTCAATGAAGTTGCATCGAAGACTTTGATTTTCA
ACGTTTATGACTTTGACAGATTTTCCAAACACGATCAAATCGGTCAAATAAAAGTACCATTGGGAGCTATAGATTTAGGT
CGAGTTATTGAAGAATGGAAGGAACTGGAATCTCCAGAAAACGATGGGGAAAAGGAAAACAGGTTGGGAGACATTTGTTT
CTCTTTAAGATACGTACCGACATCTGGAAAGCTTACAATAGTCATTTTGGAAGCAAAGAATCTTAAAAAAATGGATGTCG
GAGGATTGTCTGATCCATATGTTAAACTTTCCTTGATGCTTAATGGTAAACGAGTGAAGAAGAAGAAAACTACCATCAAA
AAGTATACACTAAATCCTTACTACAATGAATCATTTTCCTTTGAAGTACCATTCGAACAAATACAAAAAGTGAACTTGAT
CGTCACGGTTGTTGATTATGACAGAATCGGAACCAGCGAACCGATAGGTAGAATTGTTCTGGGATGTAACGCAACTGGAG
CAGAATTGCGACATTGGTCGGATATGTTGGCAAATCCCCGGCGACCAATCGCCCAGTGGCATACCCTCCAAGAGATGCCA
GAGAACAATTGATAGTGTTCGTTTGATGCTTTAAAGCGGTTTGCATAATTCGTGAAATAAGTTATCAAAATTTGAGAAAT
GTTTTTTTTTCTGAAACAAAAATGGCATGAGACAATATACCAGAGTTCATTAGTGTGTGGTTTCGTTGAATATTTTGGTG
GTGTTGCATCTTATCGTTTGTCCTCTAATCTAATAATGTTTTATTCTTTATATAGCTGATTTATTCTCGAGCGAAACATT
TTTTATGTGTACCAAAATACTTATGTCATAGCCATTGTGATTAGTCTGTAAACTATTGCGTAATAGCCCCTCTCATTGGC
TTTTTTTAAGAGTAGTGAGTTGGAAAAAAAAATACAAAAATGACTTCAATTGTTAAACATTACTCATATTTTATCAGTAA
GTTAACTTGCATATTTGCAGTTAATTATGGCATTTTATCTCTAAATCTCTGAAGCATGGCCTCAAGTCTTTTGTTTGTTT
TAATGTAATTATGTCTAGATGAGAATATTAAAATACTTCGGTTTATATAAAAGA
>SMED30030007 ID=SMED30030007|Name=Rootletin|organism=Schmidtea mediterranea sexual|type=transcript|length=6612bp
GTTGTTTTTAAATCACTATTTTTTTTATTTCATCTGATTATAATTTTTTTTTATTGAAATTTGTATTTTGAAATTGGTGA
ACAATGAGTGAGATAGATTATTCCGCTATTGTTCGGAGACGCATTGGAGATATCGATGGGGATTCAAGTGAAGATTCCAG
TTCAAAGGGTTTGCTTCAAGTTAACTTGGAACTGAGAAAGCGCTTAGAGGAAGAACACAATGAGTATCGTAGAAAGCTGA
TAGTTTTCCAAGACAGCCAACAGAAACAAGCGCAATTGGTTCAAAGATTGCAGAAAAAGGTATTGCAATACAAAGAAAAA
TGCACGGAACTTGAATCTAAAATGCAATTTGGCGATTATGAGACTCGAAAAACGAAGTATGGAAGTCCAACGAATAACGG
CTATGGGTCGCGTAGTGAGAATCCCTACGACAATCTGGATTTGGAGAGCACACTCATTAAGTTAGAAGAGGAACAAAAAA
AAACAAATTATCTAACGGAGTTCAATGAACGGCTTCAAGAACAATTAACTCAAGCAACTCAATCAAACCAGAATTTGACG
AGAGACATCAGTAAGCTTTCCGAAGAATGGCAGAGATCAAGGGAGGAGCTTGAACAGAGAGAAAAAATTTGGCGAAATGA
AGAAATGTCATTTAATGACTATTTAGCATCTGAGCATTCGAATCTGCAAAATTTGCAACGCAATACGATAAATTTCCGCC
GTCAATTTATAGAATTGAAAGCTTCAACTGAACGGGAAATAAATAATTTGAAAAATGAATTCAACCGACAGGCACGCTGT
TTGCAAACTGCTTGTTCCAATCTTTCGTGCAACATGAAATCTGCAGAATCTAAACAACAATTGAGTACTGAACGAGAAAG
TCGGGACAAGTTGAATCTCGAAAATCAACTGCGAGAAAAAAACAGAGAATTAACTGATTACCAAACTCGGTATGAATGTG
AAATGACAGAATTGCAACACCGAGTGAAAGAATTAACAGTTCTCAGTGACAAACTGAAAATTCAAAACCTGGAAAAGGAT
CGAGCATTGACAGCTTTGCAAAGAATTAAGAATGGAGATACCGCAGGAGAAATAAATATGGAAGACAAAGCTACAAGAGT
TCTTATTGAAGACACAGAAAGTATGCATCAAGCACTTCGAAATATTTCTCAAGTAGTACTAAATGATTCTGACTATTTCG
ATGCAGAAGACACAGGATTTGAAACAGTTCGAAAGAATAATCGTTCGCCAAGTCCAAGGAGAAATCGTAGCGTCAGTCCT
AACTCAAGATCAGCTAAAACACCGCCATTTGCCGATGCAACGTATGCGGCGGTTCAGTCGGCTATAAATAAAAAATCTCT
ACAGATTCAAGAGCTTAAATCAAAATTAAACAGTTCTAAAGAGAGTTGTACAAACTTCAAAAGACAAATTGATGACAATG
AAAATGAACGTCGACGATTAGAGAATCAAGTTCTCGCCATGAGAGATGAAATTGAAAGTTTACGACGAGATAAAGATGAT
AGTTTTAGAGAAGCGGATCGGTTTAAAAATGCTCTGAACATCACTTCAAATGAACGAAGCCAATTGGAAAAGATTCGATC
ATCACTAAACGATCAAATTGAGCAGCAACTCACAGAAATTGAACGACTTAAGTCAGCCAATCACGAATTACAACGATTGA
GAGAGGCATTGGAAGATGAGCAAGAAGATACTAAAAAAGACTTAGAAAGACAAATTAAAGAAAGTGAAAGATGCCAACGA
TCGATTGATGTTTTGGAAGATAAAATAAGTCGATTTAAAGAAGACGTAGTTAATTACAAAGAGGCAATTAATAAATACCA
ATTAGAAAAAGAAGTATTGGTGCAAGAAAAGGATGAGATTGGTGATGCTTTAAGTCGTTGTGAAATTTCTAAAGCCGAAT
TAGAAATCGAGGTGAATAAACGGCAAACTATTGAAGTAAATTTGGAAGAAAACATTTCAAAATTATCCTCAGAAATCGAA
GAATTGAATGATAAGCGAGCTGAAAACCACAGAACTATTAAACAGCTGGAACATGAAAGAAGTTCTCTTTTGAATGAAAA
ACGATTTTTGGAACAAGAAAAAAGTGGTTTGAAAGATGAGCTTGTTCGATCTGAACAAGAGGCTCTTGGTTTGTCAAATG
AGAAAGCCATTCAACGTCAAAGAATCGAACAACTTGAAATTAATACAGAAAGATTAGAAGAAGAGTTATCGGAACTCAAT
CGAGAAAAGCTAGATCTTACTGAACAAATAGGCTGTTTGCAGAGGCAGAAATCGGCTCTTGCAGAAGAATTGTTGCAATG
TCGTAGAGAATTAGAACGATCTAATAACAGTGTCAACAACTTATCAAAAGAAAAGGAATATCTTACCAAAGAGAAAGGAG
AACTCGTCGTTCAAGTTACAGCTTGCGAAAGGGAAAATCGTCAACAGTCAGAAGTTATTGCTTCTTTGAATTCCGATAGA
GAACAACTGGAAGCTACTCTTTATGATGCTCAACAATTAATCATTCAGTTGGAAACTCGAAAAGAGCAATTAGAAGGCGA
AAACCAAGAGTTGATTGTGAAAAAAGAAAATCTAAATGGAGAACTAGTCAGATTACGACAAGAGTTTCAACTAGAATACG
AGAAGGTTGTTAATTCACGAGAACAATTAAATTGCAGATATCAACAAATGGATAAAGAATATAAGATGGCAATTCAACAA
GAGCGAGATTCTCATGCTGAAGATGTAGAAAGACTAACTAAGATAAACGAAAGCAATCGTCATGGATCTGAAAACGAACG
AGAAGAGCTTATTCATCGATTCAATATGGAGCGTGACGACATGATTCAAAATTACGAACAAGAGAAACAAGAAATGGCCC
AGGAAATCATGACGATCCAACGAGAACGAGATGACTCATTATTAATGGCGGAAAATGAAAAGCAGCAATCATTGAGTTTA
TTAGAACAAGAAAAATGTGCAATCAATGAAAAGCTTATCAAAACGCAATTTGAACTGCAATCGGTTGAGTCGGAATTGGA
TCGCATTAAAAAGGAGTCGTCCAATCGTCAGGAGCAAGACAAACAGAACATTGCCGGCTTGTCGAGTGAACTCAAATCAT
TTAGATCTCAATTTGAGGATAAATGCCAAAATCACGAAATGGAAGTTAAAAGCCTAAATAGTGGAATCAAGGATCTAACA
TTAATGAAAGATCAAGCTCTCAAAGAAGTCCATGATTTGCATACCGGATTAAAAATAGCTGAAGAAACCCGTGATAATTT
GCGTCGGGAAATCATGGAAATGAATCGGAAATTGAAAGAAGCTGAGGAAGCCAAAGAGCACTTAAAGCGTGAAATTAATG
AATTGAAACGTGGATTATGCGAAGAACAAAGGGAAAAAGAAAATGTTATTAACTCAAATGAGAATTTGAGAAAGAAAATT
AAAGATGTTGAAACAGAAAGAATTGATTTGCACAGACAAGTAACGGATTCCAAACAAAAGGTTGCAGTACAAGATGAACA
ATTGGTCAATTCCAAACAAGAGGTCAATGAGTTAAGAAACAATCTTAAAGACGTAGAAAAAGCTCGATTAGATGTCAAAA
GAGAGTTACAAGAACTCAGACGCCATATCAAAGAGCTAGATACGGAAAGGGGCAAATTGACTAAAGAAGTCAATGATTTG
CAAACTCGTGTTGCTAGAGATGAAGAAAAAGAGGATGAAGCTCGTCGGGAAAATTTCGGTCTCAAACAAAAAGTTGTTGA
AACGGAAGCTTCAAGAGAAGCCGTTCGTAAAGATCTTGCTAATGCTCAACGACATATTGTTGATTTGGAAAATGTTCTGC
AGAGCAAAGACAGAGATTTCAGTTCCATGGTGGAATCGAGCCGCCGGGATTATAGGAAACTTGCTGATGCGAGAAATAGC
CTGCAACAAGTATATGATACGATGGTGGCCGATAATAATGAGCTGAGAATTGCTCTCAGTGGAGCAGAAGGTCGGGTCAG
TGCTCTGGAGTCACACATCGCACGAGTTGAAGGAGCTAAGAAAGAAATTGAATTCAAATTGGCTTCTATCCATAGCTCAT
TGCGAAGGCTGATAGGATTCCGACAGGAGAATGTAGTTACTATTCGGTCGAGAAACAACTCCAAATCTCCAGTTAGAGGT
CGATCAGTTAGTCCTCTTAGAAGTAGGCCGGTGTCGCCAACCAAAGGATTTGACAATACCTATGCTACTACAACTGATGG
AAAAGGTAGCCCTATCCCGCGCACAGGGTCTCCGAGTTCTAAACGAAGGCAATTATCAAGATCTCCGTCTAATTGCGATT
TCAATCCAGTCGATATAAATCCAGAAGCAGTTCGTCTGGCCTTGCGAGAATTCGTACAGCAGTTGGTTGATGCTGAACGA
GATAGAGATGACGCTTTTGCTCAAGCTCGCAGCCTAGAATCTCGATTGAAAGAACAAGAAGAACAAAATGAATATGCTGA
AAACCGTTTACAACAACTACAACAAGCTCTGTGTGATGCAGAAGAAGACAAAAAAGGCATTGATGGAAGATTGTCATCAG
CCCAAACTGCCTTAATGATGCAAGAAGAGACTATTCGGCGCAATGAGAGAGATAGAAAAGCAATGGCGGAAAAGTTGAGT
AATTACGATAGAACATTTGCATCCATGGAATCAGAAAAACGACAATTGGAAGATCGGTTGAATAAATTCCGTCATCATGA
TGCCCGATTGGAAGAAGAACATAAGCAGAGCAAAAAAGCTATTGAAAATGCTGAAAATAGAATAACTCAGCTTCAGCTTG
CCAAGCGATCGTTAGAAGGAGATCTTCAAAGAATGAAAATGGCGTTGACAGATAAAGAAACAGAAACCGAAGTTTTGGAT
GACCGCGTAAATAGTTTACTCAAACAAATACAAGATCTCGAGGGAAAATCTCAATCACTTCAACTAACGATTGACAGAAT
GAGTGTCGCCCTTGCAAAAACTGAGGAAGAAGGCAGTTTTCATAAGGACAAAATGCAACAGTTGAATATGAGTTTGAGCA
CTCATTCACAACAACTGGAACAATTACAGAATAAGATTTCTCAATTACAAAAAGCACTCGGCAGTTCAGAACAAGATCGT
CGGCTCTTAGAGGAACGATTAGAAAGCTCTAAAGGAGCATTGCATGAGGCAAAACATCAAAATAATTTGTTGATGGAAAG
AATACAAATAATTCAATCCGATTTAAATGATTCGGAAAATCGTCGATCAGAATTGGAAGGATTAAATCGACAGAGTAACA
CTTTGATTGTACAAAGACAAGAAGCTGAACAAGAACTCAATCAAAAATATCAGAAATTGAACTCGGAGAAAAACACATTA
CAAGAGCGAGTGAATAGCATGCAACGAAATTTGGCTTTGGCAGAAGCAGAAAAACAAGAAATAATAAAAAACCACGGAAA
ATTGGAAAAGGATAGATCTGCCTTGAAAAAGAGTCTTGGAAAAGTTGAACGTCAGAAATTGAAAACCGAAGAAATTGCTA
GCAAAACTGTTTTTGAACGTTCAGAATTAGATCAAATTTTAAGGAGACTTGAAGATGAAAATTGTGAATTACAGAGACAA
GTTGGAATTCTTCAAAATCAATTAGCAGAAGCTGAGCAAGCTCACTGCCACCATTTGATGGAAATCAATTCCCGCCATCG
CCAGGAAATGTCCATGGAAATGGAGCATTTGAGAGCATCGAATGTTCAAATAGAACGAACTCTAGAAGGCAGAGAACGAG
CCCACAGACAAAGAGTTAAGGGTCTGGAAGAACAGATCTCCACCCTCAAAGACCAGCTGACTCATGAAATGCAGAGGCGG
ATGGATAGTGGATATTACGAGTCTACCAGAAAGTCTAATCAATACGATTCAACTGGTGACTTGCGACATGTTCTGGATTC
CAGTTTAACTCGAGTTGCAAAAGACCCTCAACTGGACGCAATGCTACTGGAAACCGAAGCCAGGCGACTCGATGAGTCGA
TATTCATGTGTGGCAGTCGTGATCAGTCTCCGACCCGGCAGAAGCGAATCGAAATGTCAAAGAAGTCCGCAGCCCCTAAC
AATTCCAATCGGCGATTAACTGCCGGCCGTCGCAAATGAACTCGGCCTGTTCCAGTCCGTCCAGAGATGTTATGAAGCCA
CTTTTTAGATGCGTGAGTGTGAGTTCGTGTTGGTGACTGATTCATTTTTAGTAAACAGATTCTGGATTTTTGTTTTTATT
GATAGTGTAATTTCCTAATGCAGATCTGATAGTTGACTTGTTGTTCTTACGTATTCGGTATTTGTGATTCACATAGTCTG
TCATTGTTACTGTGATTTATCCTCTGTGTGACTTTTGTGATTTACTTACTTTATTGCCCCTATAATCAATAAATGGATTT
ATCGATAGTTCAGCCCTCCCCTCCAGCCCTCGTTTTGGGATCCGTAAATTTTGTTTCAATTGAGATCATTATAGTGATTT
TATTCCTTCGAAACCAAACCTTTGGGAGGGACATAAAATTAAATTATTGATTTTTTATTATCTTAGAAGTGAAAGATGTT
TATTTCCCAAATTAACTAATATCAATAATAACTTTAATTTATATCTTCATTA
>SMED30030681 ID=SMED30030681|Name=SMED30030681|organism=Schmidtea mediterranea sexual|type=transcript|length=732bp
TATGGATATACATATAATAAAAAGATTCTTCGTTTTTCTCAAGCTTCCAAGTTATAGTTTCGTAGTTTTAAAAAAAAAAT
TATTATACCCTATTACTAAACAAATAAAATTTTAATTATCTCAAAAACGAAATCCACATTTTAAAAAGCTATGTTTTTAT
AAATTTTGTGCACTTTCATTCGATCACATTTTCCATATCTCTGATAGATCGGATCGCATCTCAAGTGCAATTGATGTGCT
AATTTGTCGCATCTGTGATCGATAATTTCGAATTTTCTGTAACACCATTTTCTGATGACCCGGATACCTTGATCGAGTTT
GGCGTACAAATTATCAATCTCTATTCGACATGACTCAGTTTGAACGTCAACGCATTTGTGATTTGCGATTTCTTCCTTAT
ATTTCAAATGAAAATGAGCTCGGCGTTCATAACAGGCCTTGTGGAGAACATAAGCCTTTTCATCGCATTTCCGATCGGCT
CTGGCCCAGTCTCTGTCACATTTAGTGACAGCATTGTAGAATCGGTATTCGTGTTTGCTGCACTCAGTAACGATAAAAAG
AATAGCCAAAAATATTATGTATTTCATTTTCAAAATAAATTTTGAAAAAAAAAAAAAAAAAAGATAAATTTACCGTATTC
AGCGATATTTTTCTTTGGCATAGCATACATAATATTTTCCATTTTAAAAATGAAATAAAATGAATAAGCAAGTTGTTTTA
ATCAATCATTAG
>SMED30028196 ID=SMED30028196|Name=POU domain protein|organism=Schmidtea mediterranea sexual|type=transcript|length=1170bp
GTATTTAGTAATTTAAAAATAACCTATTTATATTATTGATTAACATAATATAAATATTTAACATCTATAAGATTTCATGA
ATTATTACTACCAATAATTTTTTTAAACTTAAAACACTTGATAAGTATCATGAAATGTTTTTTATTTCCTATTGAATCTA
CAGAAAAGGATAAGTCTTGCACCTTTGTTCTGAAAGATATTAATGAAATTCGAATTTCTGTTCCTATAAAGATATCAGCA
AATAATGATTGTAATAAAACAATTAGCTTTTCTAATTCAGATTTAGAAACTTTAGCCAAAGAAATATCAGCTGCTTTTTC
TAGTAATACTATTGAAAAGAGTACAGAAATAAAAACACTGAATGATTTTATACTTAATCAAAACAAAGATCTTCAAAATT
ATATTAATCATGTTAATGATATACCTACTTTAGATGACTGTTTGCGAAGTAAAAATTTTATATTCAAAACCGAACAAGAT
TTAGAAAGTTTGCATAAAACTCCTTCGACGGCTAGCGAGCAAGAGTCATTAAATATACTATTGAATAAATCTTTGGTTTT
TGATCTGGATTTTATTAAGCAATTTACTAAAGAATTTAAACAATTTCGACAAAATTTAGGAGTAACCCAAATGGAAGTCG
GCCAGTCCATGAAATATTATTTGAAAAGTGATTTCAGTCAAACAACGATTTGTCGATTTGAGTCTATGACGCTGGCATCA
ACAAATATGTTTGAATTGATTCCATATTTCCGAAAATGGATATCCAAAGTATATAATCCCGACAGAACCAAGACAATTAA
AAATGTCATTTCTAAACTGGAATCAATTGACATTTCTCGAGCATTGGAAGACGAGTTTGCGAAAGATAAAACGGACAGCG
AGTCCAGCAATAACAGTAATGGTGATGAAATGGTAAATGTCAAAAGCAAAACTCGGAAACAACGAATCCAATTTACGAGA
TTTCAGTTGAAAAAAATGGAAAGTCTTTTTCAAACAGTCAAACATACGCAAATTACACCGCATGTTTACAAGAAATATGC
TAAAGAATTGAAATTGAGTGAGAAACAAATCCGATGTTGGTTTACTAAGCGCAGAAGGAGTGATAGAATGCACAAGAATA
AAATGAAAATGGAAAAATTTAATTTATAAATAAAATTATGAAAATTTACG
>SMED30026364 ID=SMED30026364|Name=SMED-P53|organism=Schmidtea mediterranea sexual|type=transcript|length=2047bp
CTCATGACAGTTTTATTAAAGTTACTATTATTAGGAATTCTGCGCGTTAAACTAGACATTATCGATGGCCCAGCAATATA
TTACTTCGGCTTTCGATCCAAACTTCACAACGTTACAGCATCAGACTTCAATACATTATAAATCCTCTCCAATTGAAATG
ATTGTTCCGATGCAATGCAACCAAAACCAAGCCACCTTATCTATAACCCCTGTTCATATTTCTCCATTTGTTAATATATC
AGATTCTAATAATCACAATTTGACGAATATCGTCGAGCCTGTTTCATCAAATACTATGTCTCCTGCTTTTAAATCCGACG
ACATGCCAACATTGTCTTCTTATCCAGGTCCGTATAATTTTCAAATCTTTATTCCCAACGGAGAATTTGATGAATCCAAA
AGAAAAGGACAGACATGTGTGTTTCAAACCGATAAAATGGGAAATCACCAATTATTTACCAAACCTCATCCTCATTATTG
GAGGTTAAATTATTCAGCTGATCCTTCTATGTCAACGGAAAACATGTATATTCGGATGGTTCCAGTTTTTGGGGATCCAG
AAAAAGCTCAATGCATTTTGGAAAGATGTGCAAAACACAAAGAAGTAACAACAGATGAAAATCACTGGAAATATCGTAGC
ATGCTCATTGTAGAAAAAACCTGTGCACATTACTTTCAGGATTCGGCAACGAAAAGAGTTTGCATTTTATTACCGTTTGA
AAAGCATGCGGAAGGAGAGATTTATTCTTCCGTCAACTGTCAATTTGCATGCTACAACAGTTGCTTTAATCAAGATTCAG
GTGGTCGGAAAACACTTTATTTAATCATCACTCTAGAATTTCTCGATAAAAAAACAAATAAATTCGATGTATGGGGTCGA
CAGTGTTTGCAATTTCGCAGTTGTGCTTGTCCAAGTAGAGACTGGAGAGATAAAAAGATTAAAGGCGATCCAGAAATGTT
ACTGAAATTCAAAGAAAAACGAATCAAAACCGAAGAAAAATTAAATAATTTGGTGATTTCTAAAAGCGTCCCTATTAATA
TGGGTGGAAAGGATGCTATCATAAGAGTTCTTCCCTCGTTGCCAGGACTCGATGACGCTATTAACGCATTAGTTTGCGGA
TACTTACTGAATCGAACAACCAACATAAGCGCAATAATAGCAGCATTTAATCAGATGAAAGACGTAGAACATTTAATTAT
CGATCAATTCACATCAAATTTAGATCAAAATACATGTGACAGTAAAAGTCCTTCACAAACTCCAGAGTCTCAGATTTCTC
CGAATACATCAAACCTTCAATTCAACGATTACGGTTCACTTTACGGTGAACCATGTCAACCCTATAGACCGATGCACCAG
CAAGTTGTTAATAATTTTTCCTCTCCAGGAATTTTCAGTAAAATACCTTTTGAAACTTATCCGGTTAGTTATGACATTAA
ACTTTCACATGAAATGCCGCAGCACTTTGATGAGCTGCCATCAGACAACTATAATAGACACTGAAAAATTGCGTCCTTCC
ACGTTTAAAATTGTATAAAAAATAATATGCTTGATTTTTATTTTTGCATTTGTTTTATTTTTTAATGTTGAAAATATTTA
CGTTTGTATTATTTTTTTTTGATTAGTAGTAGTAATGCTGCTTTCAAATTTTATTGAACTTTGATCAGTGATTTTTAATA
TTCAATGACCTGAATTTTTCATGCGTCTTGATGAGGGTTTGAGTGAAAAAATTATCAATTTCTTCACTTTGTCATGTGTG
CTATGTGCATTTATTGATGTATATAACAATATATGTGTAGATAAATTTTATTTTTGTATCCGATTTAGGTTTGCAGGCAC
GATGATGAGTGGTTTTTATTTTGTAAAAATTTTTCGAGATAACATGTGCTCAAATATCAATATTATATATGGTAACATAT
TTTTGCAATAAAAATGCATTTATTGTCAATTTGGCTATAAATTAAGTATCGTACGAATCAAGCAATATTTTATCATTGCT
GTCAATATTTTGTTGATAATTTTTTATTTGCTTTGTTAATAAAGAGC
>SMED30007290 ID=SMED30007290|Name=GATA456|organism=Schmidtea mediterranea sexual|type=transcript|length=2033bp
AGGTAAGTGTCGTCATCACAAGTGGCTTTATTATTATCATTTAAATTTTTTATCATATTATCATTGATTGCTTGTTTTCC
TCATTCAAAATTTAACTGATTGTTATCAAGCTCATTTGATAACTCCAAATCTAATTGACACTCATGTAAATATCCCATTC
GAATTTCCGTTCCCGTCCGTAAGATCCACGATCCGACAATGTTTCTCCCATCGGTATTGTCGAATTCTCACCAGAGAAAC
TCACCAATAATGGACATTGTATCACCAAAATCCAACACCAGCAATCTGAATGCGTTGATCACGTATTCGAATTGTCCCAA
CATAGATCTACAGTCGAATGATCTGTCAGTGAACTTGACGAATTCTCAAAACTTCCACGAATTGGATCACAAAACCAATT
ACGGATACAAAGTTTCAAACAACGAGACAAAACTGAATGCTTTTCCATATTTAGCTCAATCGAACTTAACAGATTGCACA
ACAGACATCAGTGGAGATCTGAAAAATCCAAATTCACATCTCAACTATGGAAACGGAATTTCGTTCTCGGAATTTTCGAA
AATGTCCGAAACCAACATCCACGGATATCCGCCATATTGTTTCTCCACTAAACTGTTTCCCATTCCAAATCTACAAGAGA
AATCAATTATCAACGACAACTATTCCGTGAAATCCACCGCCGGAGGCTTTATCACCGAATCGAATGAGCATCAAAACTCT
GTAAATACTGGAAATTCCTACAATTCCAATTTAAGCTCATACAGAACGGTAATTCCCGCTCATTTTCATCTCTTCAATCA
GAATTCACCAGAATCGAACTCGTTTTACAATACGGCTTGGAACAACTATCCGTTTCAAATATCCCCAACTTATCCCATAC
AAAATCTACCGGATTCAACAATCAACTGGACAACAAACAATCAAATTCCAAGCAGCAATTTAGTTTTTGATCCCTGGAGC
TGTAAATCAAATCTTAATCTTCCTCCGATTAATGAATTCGGTTCCAATTGCGATGTCAGCAGAGAGTGTGTGAACTGTGG
AGCTAGCAATACTCAGTTGTGGTCCCGGGATAATTCTGGCTATTATTTGTGTGATGAATGCGACAGGTTCTCTCAAAACA
ACAGTAGAAACCTCGAAAAACTACCAACGAATCAAGTCTCATCAAGTACCGAAAATGAGTTCATGAAAAAATCCAATGCC
AACTTTCAACAATATGGAAAAAGATCCGACCTCGAATGCAGCAATTGTAAAATTACCAAGACTTCTTTATGGAGAAGAAA
CAACGAAGGAGAACCTGTCTGCAATGCTTGCGGCTTGTATTATAAATTACATAAGTCTCTAAGACCTCTTTCAATGAGGA
AAGAAGGAATACAAACAAGGAAAAGAAAACGGAAAACACAAATGAAACCGGTTTCCACTCATATTGCCAATAATCAAAAG
GTGCCGTCTTCAGAATCGTCATCAGAACAAAATCAGTTTATAAACACTGTGAAATCAAGTGAAAATTGCTATAACATGAT
TTCTGAAAATTCTCCAACATATATGCCAAATATATCAGTCCAGCCCTTCAATCACATTCTGAACTCCAGCCAATATCAAT
TCCAATCTACAGAAAATCCCATTAATCAATCATATTTAAATTCCTCAATCAAAAACAGCAATCCCTTTTACGATTCCGAA
AGTATACTTTATTCCTACAACAATTCAAACAAAATCAAAGATGAGCCCCTATTATCCATGGACAACGAACCGAAGCCCTA
TTCCTCAATCAACGAGAAAATTGACGTCAACTCCATAGTGAGAATTAATGAAATCTCAAACAATTCGAATTCGAGTTAAT
TATTCTGATCTACTTTTTAATTATATATTTAAGAACCGTATATACTTCGAAATATAATTTTAATGATAAAATTTTATATC
TAAAAAATGTAATTTTTAAGTGCGAATTATTAAGTATGCATTTAAATAGTGCAATTATTGTGGTAGTTATTTTTATTACT
AAATAAATGTTGTTTGTAAATTCGAGAATTTGT
>SMED30013909 ID=SMED30013909|Name=tud-1|organism=Schmidtea mediterranea sexual|type=transcript|length=2414bp
ATTTAGAAAATGGAGGCAACTTGCGATGTTTTTTTAACGAATTTACCTAAGTCGATTAATGAAGTAGTTGTGAAGAACTT
ATTTTCTTCTGCTGGGAAAGTGTTAAAAGTGAAAATAATAGATGACAAAAGAAACGAAAATCAGAAGCTATGCTTCGTTG
GATTTCGAACTATGGATGAAGCATTAGAAGCCGTCAGAATTGCAGATAAATCTGATGCGTTTGGTGATCCAATTAGAGCC
AGTATATCAAAGAAGTGTTTAGACGGAATGGAGAATAATCAAAATAGATCAAATAATAACAGAAATGATTCAAGTTATAA
AGGAACTGATTCATTTTCAAGAAAGAGTGATTCTAATTTCAAGGATAGTGAAAAGTATCCAACAAACAATGGTGACTTTG
CACAAGTAAATCGTTTAAACAACGTTGGAAGAGGAAGAGGAATCCAGACTAAAATTATTAGTGGTCCACCTTCATTCAAA
ACTAATGAAATTAAGTTAAACGATATTAAACTTAAAACAAAAGAACTGATTGACATCGAAATCGTCGATTTAAGTAGAGT
TGACATTATACTTATAAGGCCAGTAAAGTCAACGGAAAATTCAAAAATTGAAATAGCAGATTTATTGTCACAAATAAATT
CAAATAAATCGAATCTAATGCCAGTTGACTGTCCTAAAATAGGAGATTTTTGTCTTGCAAAATGGACTGAAGATGATTGC
TATTATCGGGCTGAATTGTTAGATATATTAAAGGATGGATCTTTCAAAGTTACTTTTGTTGATGAAGGAACTTCGGGTGA
TATTGTGACTGATTTGAAACACATTCCGACTGGTGCTTTACAGCAGTATCCTAAACAATTTTTGAAGGTTTCAATCGGTT
GCTCAGACGCAAAAGAATTAATGTCATTTATTTTACAAGAGTCTGAAGGAAAGGTGATTAAAATTTATTGGCTCGGAGAA
TATTGTGATTCCGTGCCTGTTGTAAATTTATTTCTGAATGAAGATGAAATTGTGGAAAAATATACCCAAATAAATAAAGT
AAATATCAGGAAAAAATTCTCACAGACATTTGATGATCTCCCAGTACATAACCTCGAAACTAATTCACATTTTTCAATAG
CACCATCAATGTATAAAAACCGAAATGATTTTATTCTTCAGATGAAAACTTTCTCGGTATCTTTAAAGCAAAAAAAATGT
CCTACGGTTTTACCAATTCCTGGTGATATATGTATATATGAGAAAAGGAATGCTAATGGCGAATCATTTCTTCGTTGTGT
TATATTAAGGCGCCAACTTTTTTCTTTTGACATCGAGTTAATAGATATCGGTGAAAAAATAATGAATGTTAATATTGATC
ACCTGTACACAGTCGAAGATATTCATGTAAAAGACGCTCCCTGTGCTATTACTTGCGATTTAGATAGTTCTATTAATATA
GATGGTATTAAGCTCCCTCAAGATTTGATCGATGGTGTTCTCTTTAATCTTAAAAAGTACAACAAAAATAATAAATACAC
TGTAAATTTCATTTTACCACAAAAGAAAGCTTATGAATATGCAAAAGTATCATATAAAGATTTGATTGGTAAAAGCTTTA
AAGTAATAATGTCGCACGTTGTCTCTCCCATTAATTTCTTTATTCAACTTGATGATGGAGAGCATCAGCTTGCACTTGGA
GATATTCAAGAAAAATTAAATATTAATTATAATGGATCACCATTCAAAAGTGCTAGCGATGTTAAAGAAACTCAGGCAGT
TATTGCTCCTTTTATTGTTTCCGGAAGTGATGCGGAACTTTTTCGGGCTTATGTTTTAGAGAAACGAAAGGAATCTCCAC
AAATTCATGTATTGTTTATTGATTATGGTAATGATGAATGGGTGAATTATGAAACTTTACTGAATACAAATGAAGAATTA
GTTTCTAAACCAGGTTTCGCAATACATTGTAAATTATCTCTAAATATTAAGAAGGAAGCGTGGAGTCACGCTATTGAATT
ATTTAATGATTTAGCTGGTTATACTGAAAATGACCAGGGGACGTTTGTAATGCAGGTTATTAGTGTCGATGGTGATTCGT
TAGTTGTGGACTTGATTTCAGAACTGAAAGGTTCGTTGACGGCCATCTTAGAAAAAAGTGGTACATGTGTAAAGTCCAAT
GAACCAATATGTAATGGAGGAGGACCCCAAGTTAAATTAAATGGCGTTTCAAATGATAAATCTCAATCAGCTATTCTTGA
AAATTTATCTGAAGGAGAATTATATAAAAAATTTATTGAACTAGAACGGAGATTAAGTGCTTTAGAGATATTGATGAAAT
AATGCTACCCGCGCAATTAGTATTATCAAATTATGATAATCTAGTCTTGATGTGTTTAAAATTTTGTATTTTAACATAAA
TTTATTCCGTGTTG
>SMED30028789 ID=SMED30028789|Name=Intermediate filament protein|organism=Schmidtea mediterranea sexual|type=transcript|length=1877bp
CTGAAAGATGGCAAGTAGAACTTCTGCAAGGTCCACAGTTACCATTAACAAATCGAGTCCTGCAGTAAAGGAATATGAAA
TGAGACAATCCTACAATTTTTCTGGAGCACCTATGGGGGGATCAGTTCAAATTCATAGCAATGTGTCTTCTGCTGTAGAA
GGCCGAGAAAGAGAAAAGAGAGAAATGCAAGATCTTAATGAAAGGCTAGCTAATTATATTGAAAAGGTAAGATTTCTAGA
AGCTCAAAACAAAAGATTAACAAATGAATTGAATACGTTACGTGAAAGATGGGGTAAAGAAGCTGAAAGGATACGAGCTT
TATATGAGATTGAAATGGATCAATTGAAAAAGTTATTAGACGAAGCTGAAGCTGCTAGATCAGAATTATTACCCAAAATT
AATAAACTTGAGGAGCAACTTCATGAATATTTACAAGGCATTGACGAAGCAAACAATCAGCATGGTATGGATCAAGAAAT
GATAGAGAAACTGAATCGACAATTGTCAGAATGCGAAGGAGAAATTGGTCTTCTTCGCCGAAGAATTGCAGCTTTAGAAG
AAGAACGAGGTCGTTCAAAGAAAACAATTAGTAGACTTAGAGATGAAATTAAACGTACAAGAATGGATCTCGATCAAGAA
ACAATGAATCATATTGCGTCCGAGAGCGAAGTGCAGTCGCGCAAAGATGAAATCGACTTTATGAAGGATTCCCATGATAA
TGAAATGAAAGAACTTGCCGCTTTAGCTTATCGTGACTCGACAGCTGAAAATAGAGATTTCTGGCGCAGTGAGATGGGCA
GTTCTCTTAGAGAACTTCAAATTGAATATGACAGAAAATTGGAAGCAATGAGAGGTGACTTAGAAAACCACCACAATGTC
AAAGTTCAAGAATTGCGACATTCATCTCCACGAGAAGGAGTCGAAAGTATTCACTTAAAAGAAGAAAATATCAGATTAAA
AAACCAATTGTCAAATCTTCGGTCAAAATGTCCGGAAATGGAGGCAAAGAATGCTCAATTAGAAAGATTTCTTGAAGAAT
TACGAAGAGAATTGGATCGTCAGATAAATGATTTTGAAAGTGAAAGAGATAAGTTGAGATCTGATTTACAACGAACCCAA
GATGAGTTAGATTCGATAATGAAAGAATTCCAAAACGTCATGGACATGAAATTGAGTTTGGAACTGGAAATTGCAGCATA
CCGAAAACTGCTCGAAGGTGAAGAATCGAGATCTGGCTTAAGAAATATGATGGAACAAACCATTTTCAAAACATCTGGAG
GAATTGAAGGAAATATTATATCAAAAACTGCAGGCAAAACACAAATGTCAAATTTGATGAAAAACGAAAGAAACGCAAAA
GTAAATCATCAAAGAACCGCCAAAGGAAATGTAATTATCCATGAAACATCTGCTGATGGAAAATTTATCATCTTGGAAAA
TACAGGAAAAAAAGATGAAGACATTAGTGGATTTAAAATCATTCGAAACATAGACAATGATCGCTCAATCATTCGGTACG
AAATACCGAACTCTACAATTTTAAAACCGGAAGGTCAACAAAAATTTCTAAAAATTTGGAGTAAAGGAAATCGACCAACT
GGATCCAATGATCTGGAATGCTCCAGTGCTAACTGGGGAACTGGAACACATATTTTTACAGCTTTGCTTAACAAATCCAA
TGAAGAAAGAGCAACTCATACTCAAAAGACCACATTTACAAATTAGCCACAATCGAAATGAAGTTTTGTGACAAAACATT
TAGAATAACTTTATAACTTATTAAGGAGATTATTTTGTTTTCTTGAAATTATATATTTATAATTCTTTTATATTTATTAT
TTCCTTGATAATAGGATTAATAAATACATATATTATC
>SMED30021661 ID=SMED30021661|Name=Carbonic anhydrase 7|organism=Schmidtea mediterranea sexual|type=transcript|length=1073bp
CATACATATATATATATATATATTGAGACTATTCATTTTTTTCCATACAAAATTCAAACAAAATGAAACCATTTCTATTT
CTATTTTTATTTCTTGTCTCATCTCTTGATCTGAAGATCACCGAGGCTTCAGGCACCGGAAATTGGGGCTATACGTCAAC
CAGTGACATCAAAGGCCCCTCGGACTGGAAAGACATTAATCCAACTTGCGACGGAGCTCGGCAATCACCGATTGATATCG
TGACAAGAAGTGCGGTTTATTTGAAAGATTTACAAAGGATTGAAATTAAAAAAATTGACAACTTTGCTAACTTTTCCACT
CTCGAAAGACATGAAACCTCTATTTACTTAAAGCTTAATAACAGTTGGGGAATCACTTTCGGAGGAAATCAATGCCTAAG
ACCTGACTCCATTCATTGGCATTGGGGATCCAAGGGTACTTTAGGATCGGAACACTCTCTAGATGGGAAGCATTATCCAA
TAGAAGGCCATCTGTTGTCATATAATTGTATGAAATACAGTTCTTTGAATGAAGCAAAATCGAGTCCGGAAGGTCTTTAT
GTAATTGGTATTTTTTATAAAATCGGAAAAAAAAATATGTTAATTGACAAAATTATCCCGGCAATGCAAAGTCTGATTTC
CAGTAGTGATATCTCGAGCGTGATCATTAACGAGATTTCGGTGGAGGAAATTCTACCGAAAAATTTGGACAAGTATTATC
AATATGACGGCTCATTGACGACCCCTCCATGTACAGAAAACGTCAGGTGGACAGTGCTAAATGAAGAGCTAACTATAGAT
GAATCTCAATTGGAAGTTTTTCGCCAATTCGACTTCGAATCTCCCAATTCTGGGGAAATGGAGAACAATTTCCGACCAAT
CTTATCAAAGTTCAATGAAAAACGAACAATATATTCCTCATTCAAAACGGGGTCCAGTTAGACAATCTTATTATGGGTGC
TTTGTGCAATCTTCATGTGCCTGAATAGTCGATAATATGATGTTCATAACACTCAAATAGCACATTTAATTCATTTTGAT
TAACACTATTTTTCATTTCATTTATGTCAATTA
>SMED30024552 ID=SMED30024552|Name=Myosin heavy chain|organism=Schmidtea mediterranea sexual|type=transcript|length=2149bp
TCACTGGAGTTAATAAACGACAAATTAAGGAAAAATGAACATGAACGATCCAGAAATGCAATACTTGGCCATTGACCGCA
AAGAATTGCTGAAAAGTTTAGATGTTGGGACTTTTGACAGTAAGAAAAATTGCTGGGTCCCCGATGAGAAGGAAATGTTT
GTGGCGGCGGAAATTAAATCCTCCTTAGGAGGTATTTTAAAAGTTCAAACGAAAAATGGAGAAAATCGTGAAGTTAAAGA
AGACGATATTCAACAAATGAATCCACCGAAATTCGAAATGATTGATGATATGGCTAATCTCACTTTTCTAAACGATGCTA
GTGTGTTACATAACCTAAAATCCCGTTATTACCGTGATTTGATATATACTTATTCTGGTCTGTTTTGTGTGGCTATAAAC
CCATATAAAAGAATTCCCATTTACACCCCAAGTATCATGCAGAAATTCAAGGGTAAACGAAAGACTGAAATGCCCCCCCA
TGTGTTCTCGGTATCCGACAACGCATATAATAACATGTTGAGAGATCGGGAAAATCAGTCCATTCTTATTACCGGTGAGA
GTGGTGCTGGTAAAACAGAAAACACTAAAAAAGTCATTTCCTATTTTGCTCAGGTGGCAGCCAATTCAAAAGATGAAGAT
AAAAGCAAAGGATCCCTTGAAGATCAAATTGTACAAGCTAATCCGGTTTTAGAAGCATATGGTAATGCAAAAACAAATAG
AAATAACAATTCCTCAAGATTTGGTAAATTCGTTAGAATTCATTTTGGATAACGGCAAAATTGCCGGAGCTGACATTGAA
TACTATCTTTTGGAAAAATCAAGAGTTGTCTCACAACAAAAAGGAGAAAGAAATTATCATATTTTCTACCAAATATTGTC
TAACAGTGCCAAAAACCTCCATGAAAAATTATTGATAGAACCAAATGCCAGCATTTATTCATTTATTAATCAAGGTGAAC
TAACAATTGACGGGGTGAATGATTCAGATGAATTCGCTTTGACTGATGAAGCGTTCGATGTTTTGGGTTTCACCACTGAA
GAAAAAATGGGACTTTTTAGATGCACAACTGCTATCATGAATATGGGTGAAATGAAATTTAAACAGCGTCCTCGTGAGGA
GCAAGCAGAAGCTGATGGTACTTCAGAAGCTGAAAAAGTTGCATTCTTACTCGGAGTAAATGCTAAAGATTTACTACAAG
CATTTTTGAAGCCAAAAGTAAAAGTTGGTACTGAATATGTTACTAAAGGACAAACTAAAGACCAGGTCTTGTTTGCTCAA
GGAGCGTTAGCTAAATCCATATATAGTCGTATGTTTTCTTGGTTGGTGACCAGAGTTAATAAAACTCTTGATACGAAAGT
GAAAAGGCAATGCTTTATTGGTGTGTTGGATATTGCCGGTTTTGAAATCTTTGAATATAACGGATTTGAACAACTGTGCA
TTAACTACACAAATGAAAGACTCCAACAATTTTTCAATCATCATATGTTCGTATTGGAACAAGAAGAGTATAAGAAAGAA
GCGATTGACTGGGAGTTTATAGACTTCGGATTAGATTTACAAGCTTGCATTGAACTTATTGAAAGACCAATGGGTATTCT
GTCAATTCTTGAGGAAGAGTGTATGTTTCCTAAAGCAACCGATGAGACTTTTGTTGCAAAATTGTATGAAAATCATCTGG
GTAAAAGTCCTAACTTTGGAAAACCAAAACCAAATGGTCAAGCAACTGCCCATTTTGAGTTACATCATTACGCTGGCTCT
GTTGGATATTCAATTACTGGATGGTTGATAAAAAATAAAGATCCAGTCAATGACAGTGTTCTTCAAACTTTAGGAGCTAG
TAAAGAAGCTCTTGTTGCTCAACTTTTTGCTCCTGCGGATGATGATGCTACTGGAAAGAAAAGGAAAGGAGGCAGCTTCT
TAACAGTATCGTTTGTTCACAGAGAATCTTTGAATAAACTTATGAAGAATTTACAAAGCACTAGTCCACATTTCATTAGA
TGTATTATTCCCAATGAATTAAAACAATCTGGATTCATTGACTCCCACTTAGTCCTACATCAACTTCATTGTAATGGTGT
GTTAGAAGGTATAAGAATATGTAGAAAAGGATTTCCAAATAGACTTATTTATTCTGAATTTAAACAAAG
>SMED30025591 ID=SMED30025591|Name=porcupine-like protein|organism=Schmidtea mediterranea sexual|type=transcript|length=3526bp
TTCCGATCTTTTCTTTTGAAATTTTTGAAATTTTAAATAAAATTAATGACAAAGGCAGAAAATCAGAGAACTACTAAAGC
ATCCGACTTCAAATTTGGAAAAGAAATCGGCCATGGCTCGTTCTCAACTGTAATGATAGCTAAGGAAATTTCTACCAATA
AACATTATGCTGCAAAAATTATTGATAAAGTTTTTATTTCTAAACAGCGAATGGTTAATACTGTTATGATTGAAAAAGAA
GTTCTCCGAAGAATAAATCACCAGTTTTTTGTATGTCTCAATTATACATTCCAAGATCCAGAGAGATTGTATTTTATTAT
ATCATATGCCGAACGAGGAGAACTTTTATCTCATTTAACAAGACTTTCGTGTTTTGATTTAACGTGCACACAATTTTATT
CCGCGGAAATTATTCTTGCATTAGAATATTTACATGGAATTAACGTTGTTCATCGAGATATAAAACCAGAAAATATTTTA
CTGAGTGAAAATATGCATATAAAAATTTGTGATTTTGGCAGTGCAATTATTATCGATAAACCAGATCTCAAGCCTTCTTC
TTTTACAGGTACAGCTCAATTCGTTTCTCCAGAAATGCTGGGATTCAGTAAAGAAACACGTGACAATAGCTATTCAGATG
AACCGAATTCTGATTCAGAGGAACATAGCGAAATTGAAGTTAAACGAACTAGTTCACCTGATAATTCTAGGGAAATATTA
TATCTAATGGACTTCTGGGCATTAGGATGCATCGTCTATCAAATGTTGTCTGGACGAATGCCATTTCACGTCAGGCATTC
ACGGCATGAATATGAAATTTTCCAAAAGATCCGAAATTTAAAATATGAGTTTCCGGTTGGGTTTGATGATACTGCTAAAG
ATTTTGTCTCTAAACTTCTTGTATTGAATCCACTGGAAAGATTAGGCAATCCATCCGATGGAAAGCACGCTGCTTTAAAA
GCCCACCCCCTTTTCCGTTCAATTGATTTCGAAATTATCGGATCACAAAAACCTCCGCTATTACTTCCAAATATTGAGCC
CATTGTTCAAAGTAATTGGGACGAAATTCCTGTTGGTTTTGAAGAAGCATGGAAATTCAATATCAAAAAAGAGTTTCAAG
AAATGGAAAAAGCCGATAAACAAAAATTATTGCAAAACCAAGAAAAACTAAATCCGTTTCATAGATTTGTACGCAATCGA
TTGATCATAAAACAAGGCATCTTGTACAAACGGCGTGGATTATTTACTCGTAAAAGAATGTTTCTATTAACTGAAGGACC
TCATTTATTTTATGTCGACCCCGATGCTATGCAATTTAAAAAAGAAGTCAGTCTAAACTTAATTACTGGCATAGAAGTAA
AAAATGACAAAGTATTTTATATCGTTACTAAAAGGCGAACATATTTTCTCGAAGATCCTTCAGCAAAGGCTGATGAATGG
CGACAGAGTATTGAATCGATTTGTGTAGAATATTATGGATCATTTAAATTATCCCCTGGCACTTGCACTCAAACGCCTCT
AGTTACATCATCTGAATCATAACTTTCGGTTTTATTTAAATTTATTTGTCATATAGAAAAAAATGTGAATTTAACGGTTT
TTAATATAATAATTTAATATTGGTTTGAATTCTTGTTTTTCATTTTAAAATTTAATGGATATAATTTAAATATTTTAATA
AATAAAAATATTCACATTGTTTTAATATTCTCTTTATGTCGTAATTGATAAATGATGCATTGTAAATGTATTTTACAACT
GATCTAAATGAAACCAATAGGAAACGAGTGGTTCGATGTCATGTGAATCAGCATGCAACCAATCATAGTTTTGTGATTCA
TTAAATATATTTTAAATTTTTAAAACAAATTAAATCACTTAATTAATTTTTAGACAATGTCACAATTTTATGGCGGATCC
ACGATATTGGCACCTCTATCGAACGTTATGGGTATACCAGTGGATCAAATTAATTTTATTGTGTGTAGTTTATTGTCAAT
CGGACTGGGTCAAATTATGCGACACAAATTCCCTCCTTCTAAATTGTCAGGAAGTACAAGAGGAATCATAGAAGCATTTT
TTGGAATACTAATTGTTCTTTTCTGCTTCGGATACCAAGTCAAGTATTTGTTACTACAATCAACAGTTTCCTATGTAATA
CTATTATTTTTGCCTTCACAAATATCAGCGCATTTGGTAACGTATTGGTGTATGGCTTTCATGGCGATAGTCCATTTATA
CAGGATGCATTACGATTACGGAGGTTATACTCTCGATATATCTGGTCCGGTTATGATTCAAACCCAAAAATTGACCAGCC
TCGCATTTAATCTTCTAGATGGACATCGAATAAATCAAGGAATAAAACTGCAACAGAGAAATCATGAATCTCACGCTGTG
AAAAAACTACCAGATGCAATCAAGTTTTTCGGTTACACATTCTACTTTCATGGTGTATTAGCGGGTCCTTTTGTATTCTA
CAATGAATATAAAGAATATATCACTGGATATGACGACAAGCATATTCCATGCTCCAAAAAGCAATTATCTTCAGCACTTT
GTTTATCAGTGATCCATGGATTATTAACAGCATATGTCGCACCGCAGTTTCCATATATATTTGTTTGCAGCACCGATTAC
CAACGGATGCATATGACTCGTAGAATAATGTTCACAATGATTTCATTTTTCCTCGTAAGACAAAAGTATTACTTTGCCTG
GTCAATGGTTGAAGCTGGTGCTCTTAGTACCGGATTGGGATATTGTGGAAAGGATAAAAACGGAGAGAATAATTGGTCGA
AAGTGAAAAATTACGATTTCATGGCAATCGAAAGCGGCAGTAGTTTGAAAGTGCTAGTGGACGGATGGAATATTTCCACA
ACAAGATGGCTCAGAGAATTAATTTATGAACGAGCTCCTTTATTCTGTCGGACAATTCTAGTTTTCACCGTCAGTGCATT
TTGGCATGGATTCTATCCTGGATATTATTTGATGTTTTTGACATTTTCTCTCTTTACGTTTTGCGCTAGAACCGTTAGGA
AACACATTCGACCATTGTTCATTGTCAGTCCAATTACACTTCAACTCTATCATGTTCTAACGAAAATATTCACAAATCTG
GCTTTAAATTACGGACAAGGACCATTTTATTTGCTGGAGTTTCAATCGGGACTTTTGTTTTGGAAGCCATATTACTTTTT
CGGACACGTTCTAGCAATGTTCTGCGTATTGACCTTGCAATTTCGGCCGATAATCAACGACAAAGTCAACATAATGAACA
ATGTTTTCACCAGAAAAGCCCAATATCGGTTTCAGAAACTAAAGATCCATTTGCAGAAAAAGTTGAGATGAAATTTTATT
GATTTTGATAATTTATATAAAATATTATAATTTTGTTTTTCGTTTTATTTTTATATCGATGAATTTATGATAAATGCTAA
ATATTCGATCCGGAATGAATTATTATTAATCTGGTTTTTATAATTGTTCATTTTTAATAGTTTTGTTTGTTTTTCGAAAA
TGGCGG
>SMED30022389 ID=SMED30022389|Name=Proliferating cell nuclear antigen|organism=Schmidtea mediterranea sexual|type=transcript|length=915bp
GTTATTAACGTTATTGGACTATAGTTTTATTATCATAAATTTTTGAATAAAATTGAGATCTTCTTTTAATAATGCTATTT
GAAGCAAAATTAAATAAAGTAGATGCATTTAAAAAGCTGATTGATTCGGTTAAAGAATTAATCCAAGAAGCAGTTTTTGA
ATGTTCATCCAAAGGTATTGAATTACAAGCAATGGATAGTTCACATGTTAGTCTAATATTTCTTTTAATTAAAAAGCAAT
CTTTTTCGTCTTATAAATTTACGAGAAATGTATCTTTGGGTGTAAGTCTTGTAAGTCTATGTAAAATACTCAAACTTTCT
CAAATTAATGATTCAATTACAATTAAAGTTGATAATGAATGTGAAACAGTCGAATTTGTTTTCGAATCATCAGATAGTGA
TCGAATGTCTGAATTCAGTTTGAAGTTAATTGACTTGGATATTGACTGTTTAGCTATTCCGGAATCAGACTATGATTGCG
TAGCAACAATCCCGTCTGTAGAATTTAACAAAATATGCCAGCAATTAGCTGTTATTGGAAGCACTGTAACAATTGTAGTA
ACTAAACAGTCAATTCAATTTTCTTGCTCCGGAGACATGGGGAAAGGGAAAATATTATTAAATCAAACTGTAAATAAACC
GAGAATAGAAGATTCTATTAATATTGAAGTTGTTAATCCATTATCAATGGTTTACTCGCTTCGATTTTTTCATATATTCT
CGAAGGCCTATTCGCTTAGCAACAGAGTTTCTTTATCTTTTGCTAATAATTTCCCAGCTTTGGTGGAGTACAGCATGAAT
GAATTGGAAGGTCATCTACGATTTTACTTAGCTCCAAAATTAGATGATGATTGATTTTGCTCTTTTTCTAGTTTTGGGGT
GATATTTTAATTGTAGTGCTTTTTTAGTATGGTTA
>SMED30006796 ID=SMED30006796|Name=tyrosinase|organism=Schmidtea mediterranea sexual|type=transcript|length=1755bp
TGTCAGTTTAGTCATGCTCGTAATCACAATAGGCATAGTCTTTGCATCTTTCTTACCTTTGAGTGACTGCCTAATCCCAA
GGATTTGCAGTAGGAATGTCACTCTTTCTAATTCAACATGTTGTCCAATTTATAATGATTTTGTTTGCGGTGGTCCTTTT
CGTGGATCTTGTGAAGTCTCCATTCTTTTTCAAAATCACACTGAGCACTCGGATGTTATCACCGGAAAATTGGAAAATGA
CGACAGACTCAACTGGCCATCAAGATTTTTTAAATTTCTTTGCAGTTGCAAGTCAAACTATGATGGAATAGCCTGTGACA
AGTGTAAATCAGGGTGGCAAGGGGAAAACTGTGAGGTGGTCTCCCCGAGAAAGATTAGACAAAATATTCTTAACATGAGA
TCATACGATGTTTTTATTTTTTTATCGATACTGAACATCAGCAAATATCAGTTGGCCGATTTCCTGATCACAAATTCGTC
CTCAAACCTTCACTGTGACCCTGTCAATGAAGACAACATTCAATTGGTCAATGCTTCCATTTATGATACAATGGTTTACT
TACATTTCTATGCGTCCAGGAGTACAATGTCTAAAGAACTTATGGAATCCTGCACAGACAAGGATAACAAATTAGACTTC
GCTCATAAAGGTCCAGCTTTTTTTACATGGCACCGACATTATTTACTGATGTTTGAAGACTTATTAAGGAGTATTGCAGT
CGAACAATTTGGATATTATAATTTCACATTACCATATTGGGACTGGACGAATCAAATAACTTGTAATATATGTACAAATG
ACTTACTTGGCGATATTCCCACTGAAGATCAAGAATCTAACAGCACATTTTTTAAATGGAAACCGATTTGCCCTCAGAAA
CCTAACGATTTTGATTGTCATATTTGCGATCCAAGCCAGGAAGAACCTTTTTTATGGAGAAAATACTCATCAACGGTAGA
CAGACTTCCTAATCAGAACGAAATTGATGAGTTATTCTCAATCAACCAATATGATTCGGAGCCCTTCAATTCCAGCTCAT
TTAGTAGGAAAAGCTTTCGGAATTTTGCAGAGGGATATTATCGAGTTCATCCAAACTCAGATGATATGAAAATGAAAGAC
ATTGGGAAGGGACTCCCCCATCAAAATATCATGATGCATAATATGTTTTATCAATGGAACTTGGATGGATTTCACAACGA
GTCCCAATGATCCATTATTTCTACTACATCATACGTTTATGGACAAAATCTTTGAAATTTGGCTATTAAGAAATAGACGA
TCTACATACCCTGGGAAATCTTCAACACCAATCGGGCAGCATGCTGACGATTTTATTGTGCCTTTCCTTCCACTAATCAG
AAACAGGAATTTGTTTACTTCAGCAAATGAATTTGGCTACGCCTACGATAACAATATTGTTGGAAATAAATTGTTCACAA
ATCATCAACTAGATTTCGAAAAGAGCTCATCGGAAATTTACTACATCCTAACGGCAGTTGCCTTTGTAGTACTAGTTGCG
ATCGTCGTCGGATTGTACTGCTTCAGACCCAGTCAAATCAGTCACACGGCTTATGAATTCATTGAAGACCAGGAGCAGGA
GGAAAAGACAAATTACTTTAACAACAATGAAAAAATGCCTCTACTGAAATGACAAGCCATTTCTGATGACTATTATAATA
TTGTCACCTTAAATTTTCTTTGAAACAACTTTATTATTGACAAATTTTGAAGAATCCTTTTAAGCAAAATAAAGT
>SMED30009513 ID=SMED30009513|Name=Glycine amidinotransferase, mitochondrial|organism=Schmidtea mediterranea sexual|type=transcript|length=1650bp
AAATCTATATAAATCTTATATGATCCTTTGATGCTTTCAAAATTCAACCGAATCCTTCCATTAAACAGTGTTTATAAAAT
TAGCGCTCCAGCAGCCAACATCAAATATAGAAATTGTTGTAGCAAATCCTCAACAGATAAACCAACAAATAAGCGTAACA
GTCCCGTATGGAGCTGGAATGAATGGGATCCTTTGGAAGAAGTAATTGTTGGTGTACCTGATCATGGTGTAGTTCCATAT
CCTTATCCAGAAACAGCGGCATGCTTTGGAGAACGTTTAACAGAAAACTTTATGAAAAAATATGGAGGAAAATACTGGAG
TGACATCATGCCTAAAGAAGTTTGGAATAACGTTTTGAAGGAACATAAAGAATTCTGTAACATCCTAAAAGGCGAAGGTG
TGACTGTGGTTCATCCAACACCAATGGATATGAAAAAAGAATACGAAACTCCATATTTCAAATCTGTCGGTTTGGATGTT
GCATTTCCAAGAGATATAATTCTCGTAGTTGGCGATGAAATTATGGAAACAACGATGACTTGGAGGTCAAGATTTTGGGA
ATATCTTTGCTACAGACCATTGATGCTAGATTATTGGAAACGGGGAGCAAATTTTCAATCAGCACCAAAACCAGTGTGTA
GCGAAGAAAGTTTTAAAAAGGATTTTGACATTTCAGGCAATAAATTCGATGAGTTGGGATATGTAACAACTGAATTCGAA
CCGATGTTTGATGCCGCAGATTTCATGAGGGCTGGCCGAGATATTTTTGTTCAACAAAGTCACGTAACAAATGCCACTGG
AATTGAATGGATGCGTCGCCATTTAGCTGAAAAAGGAATTCGGTTACACTTGTTGAATTTCAATGATCCATATGCCATGC
ATATCGATGCTACATTTTTCACAGTGAAACCCGGATACTTAATAGAGAATCCTGAAGTCATAAATCCGTATGCTGATTAT
TTGAAGAAGGCCGGTTGGAAGATTGTGAAGGGTCCAAAACCAGCAGATAGATCAGACATGTATGAAGGCATGAGTTACAA
GTGGCTATCTATGAACGTTTTAGTTCTTGATCAAAAACGAGTGTTAGTTGAAAAGCGAGAGGTTGGAATGCACAAAATGT
TTGAGTCAATCGGCATGCAACCGATCAAAGTTGATTTCGGACATTGTTTCGCTGTTGGCGGAGGGTTGCATTGTTGGACT
TCCGATGTTCGAAGAAAAGGAACTCTGGAAAATTACTTTGATTGGCCACAAGAAGTATTGGAAAAAGTATCATTGGATAA
ATAAACAATAATAGTTGAACAAGCTCCTATACTATCAGCAAAATCGACACTATCGCTGATTATATATAAATATTTATATA
TTTTGATAAAATATGTATTTATAATTTTCTCTTATTTTATTGTTATGATAATTTATAAATTTGAATTTTACTTTGTTTGA
ATCATAAACATTTATTAAAGTTATTTGTCAAAAGATGAAAATTAAAACAAAACGTGTAATCATTGCTATCTTCAATCATG
TAACAAGAAATAATTGCAATTTATCTTTATGTTGAGATAGCTCTTAGAACGCTTGGACCGATAGCGCTAAACTGTATGTT
TTATGCTTCATTTAATTACTTGAATGATAAAGGAAATTTGTTTATTGTAC
>SMED30016281 ID=SMED30016281|Name=Myosin heavy chain|organism=Schmidtea mediterranea sexual|type=transcript|length=5986bp
CTAAAATTTAGTTGATTTCGATTGTTTGAATAAAAATATCTAATGGACCCATCAGATCCTGATTTTATCTACTTGGGAGT
CGACAGGAAAGCTTTATTGAAGCAATATGGCGACTCGTTTGATAGTAAAAAGAACATGTGGATACCGGATGAAAAGGAGG
GGTACATCGCAATTACCGTTGAAGAAACAAATGGCGATAAAGTAACTGTTAAAACCGATAAAGGAGAAACTAAAGTCGTC
AAGCAAGACTCCTTGGAACAAATGAATCCCCCCAAATTCATGATGATTGAAGACATGGCGAATTTAACATTTCTCAATGA
TGCCAGTGTATTTGATAATCTGAGACAAAGATATTACAAAAATCTGATTTATACTTACTCCGGACTGTTCTGTATTGCAG
TGAATCCTTATCGAAGATTTCCGATTTATACGTCCCAAGTAATTATGAAATATAAAGGAAAACGTAGAGGAGAAATGCCA
CCGCATATTTTCTCAATTTCGGATAATGCTTACTCAAATATGCTTGTCGATCGAGAAAATCAGTCTGTTTTGATTACTGG
AGAAAGTGGTGCTGGGAAAACTGAAAACACGAAAAAAGTTATTACTTACTTCGCCAATGTAGCAGCAGCAAGTAAAAAAG
ATGATGACGATCCATCGAAGAAAAAAGGTTCATTGGAAGATCAAATTGTACAAGCCAATCCTGTACTGGAGGCATACGGA
AACGCCAAAACAACTAGAAATAATAATTCGTCAAGATTTGGAAAATTCATCCGAATTCATTTTGGCACTAGTGGAAAAAT
TGCAGGAGCTGATATTGAGTTTTATTTGTTGGAGAAATCTCGTGTTATCTCACAGCAAAAAGGAGAAAGAAATTACCATA
TTTTCTATCAATTGCTTTCGGCAGGAGGGAAAAGTTACCACGATAAACTTTTGGTTACTGCTGATCCTGCTCTGTTTTCT
TTTATCAACCAAGGAGAATTGACGATTGACGGTGTTGATGATGAGGAAGAAATGAAACTTACTGATGAAGCCTTTAGTAT
TCTTGGGTTTTCCGATGAAGAGAAAATGAGTTTGTTTAAATGCACTACTTCAATCATGAATATGGGAGAAATGAAATTTA
AACAAAGACCCCGTGAAGAACAAGCCGAGGCTGATGGTACAACAGAAGCTGAAAAAGTCTCTTTCTTACTTGGAGTGAAT
GCCAAAGATCTAATGCAAGCAATTCTTAAACCCAAAGTCAAAGTAGGAAACGAATATGTAACGAAAGGTCAAAGCAAAGA
TCAGGTTTTATATTCTCTTGGAGCCTTAGCAAAATCTTTATATAACAGAATGTTTGCTTGGTTGGTTACTCGCGTGAACA
AAACGCTGGACACAAAAGTCAAAAGACAGTTTTTCATTGGTGTCTTAGATATTGCCGGATTTGAAATATTTGATTTCAAT
GGTTTCGAACAAATTTGTATCAACTATACTAATGAGAGACTCCAACAATTTTTCAATCACCATATGTTTGTTCTTGAACA
AGAGGAATATAAAAAAGAGGCCATTGACTGGGAGTTCATAGATTTTGGAATGGATCTCCAAGCTTGCATTGAGTTGATTG
AAAAGCCTATGGGTATTTTGTCGATTTTAGAGGAAGAATGTATGTTTCCCAAAGCAAGCGATCAAACATTGAAAGCTAAA
TTGTACGATAATCACTTGGGTAAAAGTCCGAATTTCGGAAAACCAAAACCACCAAAACCAGGACAACACGAAGCCCATTT
CGAATTACATCATTATGCTGGCAGTGTTCCATATAATATTACTGGGTGGTTGGAGAAAAATAAGGATCCATTGAACGAAA
GTGTTGTTGGATTACTCGGAGCATCTAAAGAATGTCTAGTAGCTTCTTTATTTGCACCGGCTGAAGATACAGGAAAACCC
GGAACCAAGAAAAAGGGTGGATCCTTTATGACTGTGTCATATATGCACAGGGAATCATTGAATAAATTGATGAAAAATCT
AATGAGTACAAGTCCACATTTTATTCGTTGCATTGTCCCCAATGAATTCAAACAACCAGGAGTTGTTGATGCGCATTTGG
TTATGCACCAACTTCATTGCAACGGTGTATTGGAAGGTATTAGGATTTGCAGAAAAGGATTCCCAAATAGAATGATCTAT
TCTGAATTCAAACAAAGATATTCAATTTTGGCACCGAATGCCATTCCACAAGGATTTGTTGAAGGCAAACAGGTCACTGA
TAAAATCCTCACAGCCATTCAACTGGACACAAATCTTTATCGCTTAGGAAATACCAAAGTATTTTTTAAAGCTGGAACAT
TGGCTCAACTTGAAGATATAAGAGACGAGAAACTTTCGTCGTTGATATCGTTGTTTCAAGCTCAAATTAGGGGATATTTA
ATGAGACGGCAATACAAAAAACTTCAAGATCAAAGAGTTGCTCTGTCTTTGATTCAAAGAAATATTAGAAAATATTTGGT
TTTGAGAACATGGACCTGGTGGAAATTGTTCACTAAAGTAAAGCCTATGCTTAATATTGCTCGCCAAGAAGAAGAAATGA
AAAAGGCAGCCGAAGAATTAGAAAAGCTGAAAGAGGAATTTGAAAAAACTGAAAAATATAAGAAAGAACTTGAAGAACAA
AATGTTACTTTACTTCAAGCAAAGAACGATTTATTCTTGCAGTTGCAAACTGAACAAGATAGTTTAGCTGACGCTGAAGA
GAAAGTATCTAAATTAGTTCTACAAAAGGCTGACATGGAAGGAAGAATTAAAGAACTTGAGGACTCACTTTCAGAAGAAG
AAAATTCAGCTTCAACATTAGAAGAAGCAAAAAAGAAACTCAATGCTGACATCGATGAATTGAAGAAAGATGTTGAAGAT
TTGGAATCATCTTTACAAAAAGCAGAACAAGAAAAAGTTGCAAAGGATCAGCAGATTAAATCCCTCAACGACAATGTTCG
TGAAAAAGAAGAACAATTAGCCAAAATGCAAAAGGAAAAGAAAGCTGCTGATGAATTACAAAAAAAAACAGAAGAATCAT
TGAAAGCCGAAGAAGAGAAAGTCAGTAACCTCAATAAAGCTAAAGCTAAATTAGAACAAGCCCTGGATGAAATGGAAGAA
AATCTTGGTCGAGAACAAAAAATTAGAGCAGATGTTGAAAAAGCAAAACGAAAAGTTGAAGGTGAACTTAAACAGAATCA
AGAGTTATTAAATGACTTGGAGCGCATAAAATCCGAACTTGAGGAACAATTAAAACGAAAAGAAATTGAATTGAATGGTG
CTAACTCTAAAATCGAAGATGAATCTAATTTAGTTGCAACACTTCAACGGAAGATCAAAGAACTTCAAGCCAGAATCCAA
GAATTAGAAGAAGACCTTGAGGCGGAAAGACAAGCAAGGGCAAAAGCTGAGAAAGCTAAACATCAATTGGAAGCGGAACT
TGAGGAAATCAGTGAAAGACTAGAAGAACAAGGAGGTGCAACTCAAGCCCAAACAGATTTAAATAAAAAACGAGAAGCTG
AACTCATAAAATTAAAGCGAGACTTGGAAGAGGCTAATATGCAACACGAACAAGCCTTGGTGCAAATGAGAAAGAAACAA
CAAGATACTAGCAATGAATTCGCAGATCAGTTAGATCAATTGCAAAAATCAAAATCAAAGATTGAAAGAGAAAAGAGTGA
ACTTAGAGCAGACATTGAAGATTTATCAAGTCAGCTGGAATCACTGAATAAGACCAAGGGAAATCTTGAAAAGTCAAATA
AAGCATTAGAAACCACAGTTTCAGATCTTCAAAACAAACTCGATGAATTAAGCAGACAATTGACAGAAGCAGGAAACGTC
AACAATCGAAATCAACATGAAAATAGTGAAATGCACAAAACCTTAGAAGACGCCGAATCACAAATAAACCAACTCGGAAA
GGCTAAACAACAACTCCAAGCGCAGCTAGAAGAGGCTAAACAAAATTTGGAAGATGAATCACGCGCAAAAGCAAAACTTA
ATGGAGATTTAAGAAATGCGATGAGTGATTTAGATGCATTACGGGAATCATTGGAAGAAGAACAAGAAGGCAAAAGCGAT
GTTCAAAGACAGCTTGTGAAAGTCCAAAATGAATTACAGCAACTTAAAGCAAATTCTCAAGGTACTGGAGGAGTGAGAAC
AGAAGAAATGGAAGAATTCAGACGAAAGATGAATGCCAGAATCCAAGAACTAGAAGAAGAATCGGAATCAAATAAATCTA
AATGCAGTCAATTAGAAAAAATGAAATCCAGACTTCAAGGAGAAATTGAAGATTTAGTGATAGATGTTGAAAGAGCCAAT
GGATTAGCATCGCAACTGGAAAAGAAACAGAAAAATGTTGACAAATTAATAGGCGAATGGCAAAAAAAATATTCAGAAAG
TCAACAGGAACTAGAAGTATCACTGAGAGAATCAAGAACAGTTTCCGCTGAAGTTTTCAAATTGAAATCACAATTAGAAA
ATTCTCAAGATCAAATCGAAAGTTTAAGAAGAGAAAATAAAAATCTATCTGATGAAATTCATGACTTAACTGAACAATTG
GGAGAAGGAGGACGAAATGTACATGAAATTGAAAAGACTCGAAAAAGAATTGAAATTGAAAGAGATGAATTACAACATGC
ATTAGAAGAAGCTGAATCAGCTTTGGAACAAGAAGAAGCTAAAAGCCAAAGAGCTCAGTTAGAAATAAGTCAAGCCCGTC
AAGAATCTGAAAGAAGACTTGCAGAAAAAGAAGAAGAATTTGAAGGTACACGAAAAAATCATCAACGAACACTCGAATCC
ATGCAAGCATCTCTTGAAGCTGAAGTTAGAGGACGAACTGAAGCAATGAAAATGAAAAAGAAGCTGGAGCATGATATTAA
TGAGCTTGAAGTTGGGTTAGACACGGCTAATAGACTGAAATCTGAACAAGAGAAAAATGCTAAGAAATATCAGCAACAAT
TAGCAGAAATGCAAGCTCAAGTTGAAGAACAGCATCATCAACGAGAGCAAGTTAGTGAACAGCTTGCGTTAACAGAACGA
AAATTAGGTATGATGCACGGAGAGCTGGAGGAAATGAAAAATATGTGTGATCATGCTGAAAAAAATAGAAAAGTCGCTGA
ATCTGAAAAGAATGAAGCCATTGATAAACTAAATGAGTTTTCAATTCAAAATTCTTCATTCATGGCAACTAAAAGAAAGA
TGGAAGCAGATTTAGCAGCAATGCAAGCTGACTTAGAAGAATCTACAAATGCAGCTAGACAAGCTAATGAACAATTGAAA
AAGGCAATATTTGATAATACTCGACTTTTCGATGAAATTAAACAAGAACAAGAACATGCTCAACAAGCTGAAAAAGCAAG
AAAGAATTTGGAGAGCCAGTTGAAAGAATTGCAATCAAAACTTGAAGAAGCAGAAGCAAATGTACTTAAAGGAGGCAAAA
AGGCTTTGAGTAAATTAGAGCAAAGAATAAGGGAATTGGAAGGCGAACTAGATGGTGAACAAAAACGACATGTTGAAACT
CAAAAGAATGCTAGAAAGTCAGATAGGCGACTGAAAGAAATTTCGTACCAGATCGACGAGGATAAAAAGAATCAAGATCG
AATGCAACAACTTATTGAAAGTTTACAAGCAAAAATAAAGACTTATAAACGACAAGTAGAAGAAGCTGAAGAAATCGCTG
CTGTGAATTTGGCAAAATATCGCAAAATTCAACAAGAGATTGAAGATTCTGAAGAGCGAGCTGATCAAGCTGAACAAGCT
TTACAAAAATTAAGAGCAAAGAATAGAAGCTCGGTATCTGTTAGTCGAGGAGCAAGTTCTGGACCTGGAACCAAAATTTC
TATAACTGCGACATCAATCCGCAGTGAGGACTAATCTGACAGAAAAAATTCATCATTAGGAAATATGCAATTAGTTAGCC
AATATTTCCCTTAGACTAGATTTTAAACCTTTAAGACATTTTTTTCAAATTAATAAAACGTTTTGA
>SMED30001958 ID=SMED30001958|Name=POU domain protein|organism=Schmidtea mediterranea sexual|type=transcript|length=1820bp
GAAGATTTCAACCAATTTCATTGTTAAAATTTTCATAAAATCGCCGTAATTGGCTGTCCGTTGCGGCAGTGGCAGTCAGG
CAACTCCGTTCACTCAGCAGCCAATAAACTCGTTTGGAAATATTCCTACTCAAATCAAATTGATTTCTCATAAAAAAATT
TGCATATCTTCTCACTTTTTCTATATGCATTTCATTGTTACTTGGTGAAAACTGAATGCCATGACCCAAGTTAGTCACAT
AGGTTCCTCTCATTTCAATATTAATTCTTCAATCAAAACTGCTCTTCCTTTAATTAATTCTCAATACAATTCATTGGTAA
CCGAAACATCTATTGGTTCCAACATCAACCTACCGTTTCGACTCGATGCTCCATCTCTATTCAATGGGCTACCGGAAGAA
TGGGGTCCCACTGACAGCGAGTTAACTCAAGCGGTCAATCATGTAAATTACGCCGGATTATCCGAGTCGAGCATTGATTA
CAAGTTTTGCAGCACTCTCAATTCTTATTTTGACTCAAATCAAAGATTTGGAAATGTACCAAACGACACGTATTTAATGG
CAAATTCCATGGCAGCCAACATTGAGAACAATAGCTGGCATAACAGCCGATATTACTTCGACAGTAATCATCCTCTCAGC
CAATGGCTCCCGGTGCCCGTTAACTGTTGGTCCAATAACAATTACATTCGCGATGACTTTTACGAGAAATCTCTCCAGGA
AGATGAGTTCAAATCCCGCTCGGTAATACAAGCAACTGAGAAGAATTCCAATCCGGTAAATCCCGACCTGAATCCTTACA
AATCTATCATTTCCGATGAATCAGAAAGCATGAAAATGGCTAAAACCTCCCACAACCTTATAAATAATGACTCGATGAAT
ATGTGTAGATTTCCTCAAAATGATATTATTATGCCAGATTTAATAAATGCCCCAAGTCCATCACTGACAAATAATAATAA
TAATAATAAAGTAATAAATAATTATTATATAATGAAAATTAATATATTAATAAAAAATATTTATGCACGGTCTCAATTTA
CATCTAAATGCCATCAAATTTGGCAGGGATACTTCTAATATAAAACCTAGATCAGCTGAAAGACTTTTATATTAGATAAA
TAATCGTTAATAAATAAAACCCAATTTGCGATAAATTATGTGTGGCCTCCAGGCGGTTCCTATCAAATAAACATATGTAA
TTATACGTGTAAACACGTGCATTAAATATCTTCACAAAGTCAAATGAGTTGATTGGACAGAAAAATTAGAATCGGAACTA
ATTCTGCAATCATTCAAAAATTTGAGCTAATCTTTATTCTTTAAAAAGCAGAGAATCAGCGGTTCTTTACATTCATGCCA
GTAATGAAAAAAGTAAACCGCAAATGACTTTTATTACTGTTCTAGTAAATGCATCCCCATAAGGAGAGTAATTCACATAA
AACTTTAATTGTCAATAAAATTATTACTGGGACATCAGCGCATCCTGGCATGCACCGCAGCAATGAAAAAGTAGACCGAG
AAACTCTGAGATTATTGAGCCATCAAACGTATCACTCATTTTCTTATGTTTGAATAAAATATTTACAAATTTTAATACAG
AGATAATCGATGGAGGAGGAACACTACTATTTTTGCTATCTATAGCAATCAATAATTATGTGAAAATATAGCTGATCAAA
ACAAGATGCATTAAAACAATTTTACTTCCTATTATATACGTTTTTGAATCCTGAGTGAAGCCAGTCGGGTTAACGAATAT
TATTAAAATGAAAATAACTATAAATCAATAACTATACCTCATCACATGATGAATGGTCCA
>SMED30019471 ID=SMED30019471|Name=Synaptotagmin|organism=Schmidtea mediterranea sexual|type=transcript|length=1910bp
TTTGGGACTTCTCCAATCTATTAAATATATTTAGAACGAACATCGTTGTGAGAAGCTTTGTCTTGCAAATGCACACCAAC
ACATTCTCTGCCTACTCGAGACCAGCCTGAAGGACGCACCCAGTCATCTCTTATAAAAAAAAAACTCTTCTCTGATTCGG
TATTTTCTGATTAATGGAGATCAAACAATGATTGGGCAACGTCTAGTTTAGAATATTTACAACTAAATCTGCGTATTTCA
ATTTTTTTTCGATTTTGATTGTTGCAAATATTTATGAGTATATCCAATAATATAAGCAGCGCTAGTAATATGACGATAAA
AATGCAACAAAATGTTGAAAAGTCTGTATCACAGCATGTAACAGCGCAAACAAAGGAATCAACATCAAATAAACTTCATA
CAACAACAGCCGGTATCTCGGTTTTGGAGAAAACATTAAAAACATTTCACATGAGTACTGGCACTTTGATTGGAATCGGA
GTTACAGTGTCTATCGTAGTTTTATTACTTCTTTATTGTATTTGCAAACGATGCATTAAGAAACGAAAGAAAAAGAAAGA
TGGCGGCAAAAAAGGTGGAAAAGGCGTAGACATGAAAAGCGTACAACTATTAGGAAATGCTTATAAAGAAAAGGTACAGC
CGGACCTAGAAGAACTGGATGCTAACATGGAAGATAATGAAGGGGCGCAAGATAAAGAAGCAGTCAATCTTGGAAAATTA
CAGTATTCCCTTGATTACGATTTTACAAAAGGCGAGCTAACAGTCGGAGTAATTCAAGCCACAGATCTCCCAGCAATGGA
CATGTCAGGAACTTCTGATCCATATGTGAAAGTTTTTCTATTACCGGATAAAAAGAAAAAATTCGAAACCAAAGTTCATC
GAAAAATATTAAATCCAGTCTTTAATGAGACATTTGTATTCAAGATTCCTTATTCTGAGGTCGGGTCTAAAACTTTGGTA
TTTAATGTTTTCGATTTCGACAGATTTTCAAAGCATGATCAAATTGGGCAAATCCAAATTCCACTGGGTTCAGTAGATTT
ATGCAAAGTTATTGAGGAATGGAAGGAACTCGGCCCACCTGACAGTGATAATGAAAAGGAAAATCGATTAGGAGACATTT
GTTTTTCTTTGAGATATGTTCCAACGTCTGGAAAGTTGAATATCGTCATTCTGGAAGCTAAAAACCTCAAGAAAATGGAT
GTTGGAGGATTGTCGGATCCTTACGTGAAACTATCGTTGATGCTTAACGGCAAACGAGTGAAAAAGAAGAAAACCACAAT
AAAAAAATACACATTAAATCCTTATTATAATGAGTCATTTTCATTCGAAGTTCCATTCGAGCAAATACAAAAAGTTAATT
TAATAGTGACAGTAGTTGATTATGATAGAATTGGAACAAGTGAACCTATTGGAAGAGTAGTTCTTGGATGCAATGCTACA
GGAGCAGAACTTCGGCACTGGTCCGATATGCTTGCTAATCCTAGGAGACCAATTGCCCAGTGGCATACCCTTCAAGATAT
GCCGGAGAAGAGCTGATTTCATTGATTTTAATTTTAAAATCTGTCATATTTTGTGATAAATTTTGCGTATTTATACAATC
ATTCGATTCGATCTTTCCATCAAGTTTCGTAAAATCTGCAGACATTCACGAGGATGCAGAAGAAATAGAAACTTTGTAAT
AAATTAGCATTAGCTGTGATCTCCTATTGACCAAAAAACATAAACTTAATTTTTATAAAGAATTGTGGTGAAATGTTGTT
AGGTTTCAGTCATAGATAACTGTATTTGTGTAATAGTTTGTGTATTTCTCGTCCTGGCCCGATTTTGTGCATTCCACTGG
CAATACTATTTCACTAATAAATTATGTCTGATAATGTATACCAGTATAGGATTTTGTTTGAATTAATGAA
>SMED30022992 ID=SMED30022992|Name=Myosin heavy chain|organism=Schmidtea mediterranea sexual|type=transcript|length=6157bp
ATATATATGCATCTCAAAATTGGAAATTAAATTATTAATTTTAATTTACTAACAAATTTTTGTTGTAAAATGGAAAATAT
TGATGCTAAAGACTTGCAATATTTACAAGTTGATAAAAGTTTAGTTTCTGACTCTGCTAATTTGTCTGATTGGGCTAACA
AAAAATTGGTTTGGATACCTGACGACAATGAAGGATTTATTTCAGGATCACTTATAGAAGAAAAAGGCGATGAAGCTACT
GTCAAACTGGAAAATGGCAAAACAATGAAATTACCAATTGAAAATGTTCAAAAAGTGAATCCACCAAAATTTTTTAAATC
TGAAGATATGGCAGATTTAACTTATTTGAATGAAGCATCAGTTCTATATAATTTAAAAGACAGATATTTTTCAGACTTAA
TATATACATATTCCGGATTATTTTGTGTCGTTGTAAATCCATACAAGCGCTTGCCTATTTATACAAATAATGTAATTGAG
TGGTACAAAGGTCGAAAACGTCACGAAAGACCGCCACATATTTACGCAGTTACTGATGTCGCATATCGAAATATGCTCCA
AGATAAAGAAAATCAATCAATTCTTTGCACTGGTGAATCCGGAGCAGGAAAAACTGAAAATACGAAAAAAGTTATTCAAT
ATTTGGCCTCAGTTGCAACTTCTTTGAAAAATCAAAAAACCACAGCCAGTAATGCATTGTCACAATTTTATGCTCACGAT
GTTAACATTGGAGAATTGGAGACCCAATTACTTCAGGCCAATCCGGTTTTAGAAGCTTTTGGAAATGCAAAGACTATCAA
AAACGATAATTCATCTCGTTTTGGTAAATTTATTCGAATCAATTTTGACACTTCTGGATTCATTTCCAGTGCAAACATAG
AAACATATTTATTGGAAAAAGCTAGAGTTATTAGACAGGCAGCTAATGAACGGTGCTTTCACGTGTTTTATCAGCTTCTT
CTTGGTGCAGACGACCGATTGAAAAAGGAATTAATCTTAGAAAATGTTTCTACATACAAACTTTTATCCAACGGCATGAT
AGCTGTACCAGATTACGATGAAAAGCAAATGTTTAAAGATACAGTCGAAGCTCTCGATATAATGGGGATTTCGAAAGACG
AACAAGAATCTATTTTCCGAGTTATAGCCGCAGTTTTACACATGGGAAATATCGAATTCAAACAAGAAAGAAGTTCGGAT
CAGGCTGCTTTACCAGACAATACAGTTGCTCAAAAGGTTGCTCATCTTCTCGGTTTACCGGTCACAGAAATGACCAAAGC
ACTGCTGAAACCAAAATTGAAAATTGGAAGAGAAGTTGTAGCAAAAGCTCAAACTAAAGAACAAGCCGAATTTTCCGTTG
AAGCTATAAGCAAATCAACATATGAAAGAATGTTTCGATGGCTGGTAATGAGAATAAATAGATCAATTGACAGAAATAGA
CAAAAAACTAATTTCATTGGCATTCTTGATATTGCTGGCTTTGAAATATTTGAAATAAACTCTTTTGAACAATTGTGTAT
AAACTATACCAACGAGAAATTGCAACAGCTTTTCAATCATACAATGTTTGTGCTAGAACAAGAAGAATACAGAAAAGAAA
ATATCAAATGGGAATTCATTGACTTTGGTCTTGATTTGCAGCCAACTATCGAACTAATTGAAAAGCCAATGGGTATTCTT
TCTTTGTTAGATGAAGAGTGCTTTTTCCCAAAAGCTACCACAAAGACATTTGTTGAAAAATTAATTAAAAACCAATCGAC
GCACCCTAAATTGAAAGCTACTGATTTTAGAGCAAAAGCAGATTTTGGGGTTATTCATTACGCGGGAAAAGTTGATTATG
TCTCCGATAATTGGCTGGTGAAAAATATGGATCCTCTTAATGAGAACGTTGTTAGTCTCTTACAAGAATCAAATGAAAGT
TTTGTCCAGACAATTTGGAAAGATGCAGAAAACATTATTGGCTTATCAACTACTACCGCCCAAGAATCTGCCTTTGGAGG
TGCAAAAACTACTCGCAAAGGAATGTTTAGAACCGTCGGTCAGCTTTACAAAGAATCATTAACGAAATTGATGGATGTTC
TGAATAATACCAATCCCAATTTTGTGAGATGTATCATTCCCAATCATGAAAAAAAATCTGGTAAAATCGATTCAAAATTG
GTTATTGATCAATTAAAATGTAATGGTGTGCTGGAGGGTATCAGAATATGCCGACAAGGATTTCCAAACCGAATTTTATT
CCAAGAGTTCAAACAACGTTATGGAATTCTTACACCAAATGTCATACTAAAAGGTTTTATGGACGGTCGAAAAGCAACAG
AATTAATGCTTGGAGAGTTGGATATTGATGCATCAAATTACAGAATTGGTCAGAGTAAAATATTTTTCAAAGCTGGAGTT
TTGGCTAGATTAGAGGAAGATCGAGATATTAAGTTGACTGAAATCATAGTCAAATTTCAATCTTATGCACGTGGATATTT
GGCAAGAAAAAATTTACAAAATCGATCGCAAAATCTCAATGCAATTAAAATTATTCAAAGAAATTGTAATGCATACTTAA
AATTGAGAAATTGGGCCTGGTGGAAATTATTCACACGTGTTAAGCCTCTATTAAGTGTTACAAGGCAAGAGGAAGTTGTG
GCTTTGAAGGAAGAAGAATTGAAAAAATCAAAAGAAACTCTCGAAAAGATGACTGCTGATTTCGAAACAACTAAGAAAAA
TTATGAATTTCTCATGGAAGAAAAAAATAAAATTCAGGAAGATTTGGAAAAGGAGCGTTTTGCATTGCAAGATATGGAAG
GGGAAAGAGAAAATTTGATCAAGCGATTAAAAGATATTGAATCAGATGCTCAAGAATGCGAGTCAAGAGAGATGGAAACA
AAAGACAAATGCAATAAATTGGAAATCGAGAAAAATAAATTACAAAATGAAATTTCACAGCTTAGCCAAAATTTAGAATC
AGAGGAACAGAACTTGCAAAAAGCTCAAGCAGACAAATTAAGTGCTGATAAACGAATTAAGGAACTGGAACAAAAAGTGG
CCGAATTAGAAGATAAACTTAACAAGTTAACTAAAGAGAAAAAGTCTCTAGAAGAGCGATTGGCTGATGTTATGTCTACT
TTGGCGGAAGAAGAAGAAAAGAGTAAACAATTAACGAAACTCAAATCCCGTCACGAAACAAGTATCACTGAATTAGAAGA
GAGATTGAGCAGAGAACAAGCAGCTCGACAAGATTTAGAAAAAACAAAGAGACGACTAGAAACTGAATTGTCTGAAAAAT
CTGATAATTTATCATCTCAAAGCCATACATATGAAGAAATTAGAATTACTCTTGAACGAGCAGAAACACAAATAATTCAA
ATGCAAGCTAAATTAGATGAAGAAACAACGGCAAAATCGTTGGCTCAGAGACAAATTCGAGACGCAGAAAACGTAATCAC
CGAAATCAAAGAAGATCTTGAAGCTGAAAAACGTTCAAAAGAGAGAGCTGAAAAAGCTAAGAGAGATTTAGCCGAAGAAA
TAGAATCTCTTAAAATGGAATTACTCGATACGGGAAATACATCAGAAGAACAGCAAAGCGTATTGCGCAAAAAAGAAGTT
GAATTACAAAATTTTAAAAAATCTTTTGAAGATGAAACAAAACGAACTGAACTTGAAATGCAAGAAATTCGTAAAAAAGC
CGCAGTAACGATTGAAAACCTCAACAATCAAATAGAAGCTGCTAACAAATCAAAACTTGTTGTTGAAAAGGCTAAATTAC
AGTTGCAAGAAGAACTTAATGATGCAAATAACGAATTAGCAAATTTAAGATCCTCCAAAGCTGACTCTGAAAAACTTAAG
AAGAACACAGAACTACAACTTTCAGAAATATCTCATCGCCTTTCAGAATCTGAAGAACACAAGCTAGATGCTGAATCAAA
ACTTAAAAAACAAACCGAGTTGGCTGACAAATTGGCTATGAATCTCGAAGAAATTGAAAGCAAATTGTCGCAATTAACTA
AATCTGAATTATCAGTTCGAAGTCAGTTAGCAGATTTACAAAAGGACTTTGAAGAAGAAACTCGATTAAAATTGAATGCT
CAAACTACATTAAGGCAGTTAGAAAGTCAATTTGCTTTATTAAAAGATCAATTAGAAGAGGAGCAACAATCAAAAGAAAA
TTTAGAAAGTCACGTACAAGCTCTACAGAGTCAAATGCAAGATGTCAAAAAGAAAGTTGAAGAAGATGCTCAACAGCTAT
TAGAATTTGATGAATCTCGTAAAAAAGTAATGCGTGAACGAGAAGAATTGCAGATTAGACTTGAAGAAGCAATCAATCAG
ACTGAAAGATCAGAAAAAGCTAAGCGAACAATTCAAGCCGAGCTTCAAGATGCATATCATGCCCTAGAAAGTCAGAAAAA
TGACCAAACTCAAAGTGATCGAAAACTTAAAAAAATGGAATCACAAGTCAATGAAACATTGAACTTGAATAAAAAATATT
TAGCTGAAAAGGATACATTAGAAAAAGAAAATCGTGAAAAAGAAACAAAAATCATGCAATTACAGAGAGAAATGAATGCA
AATGACGATAAACTTTCGGAATTGGAAAATGCAAAATCCTTACTGACTCGGCAATTGAATGAATTGGTTTCGAGCAAAGA
TGATGTTGGTAAAAATGTTGGCAGTTTAGAAAAGGCTAAAATTCAATTAGAAACCGATTTAGAGAATTGTAAAGTTACTA
TTGAAGAATTAGAAGAAGAAAATGCAACTCTTCAAATGAATAAAGAACGAGTTGAAATGCAATTAACTGCTACTAAAGCC
CAACTGGATAGGGAGCTAGCTTCCAAAGATGAAATGCACGATGAACAGTTCAGAATTATAAACAAAAAGCTTCGTGATGC
AGAAGCGGCACTTGAAGAAGAGCAGAAATCCAAAACAGCTATGGTTTCATTGAAAAAGAAAATAGAAAATGAGAACATGG
ATTTATTAAATCAATTAACAGAATCTGAGAAGGTCAAAGAAGAGCTGGCCAGACAGATTAAAAAATACCAAGGAAATTCT
AGCAATCTTCAAAGAGAAATGCAAGAACTTGTTGCTGCTAAAAATGATTATTTGAAGCAGATTAAAGAACTAGAAAAGCG
CTTACGTACTTCTGAAAGCAATGTAAACCAATTATCCGAAGACCTTTCGCAGTCTGAACGTGCTAAGAGAACCATTGCAT
CTGAAAGAGATGAAGCTTTAGAAGAAATTGCCACAATCACTATGGCTAGGGATACAGCTATTGCAGATAAAAAACGTATT
GAGAGTAGCATTTCAGCATTAAGCATGGATAATGAAGAATTGCAATTACTTTTGTCAGAATTAGAAGACAAAAATAAGAA
AATTATTTCTCAGTTAGAACAAGTTCAAAATGAATTAAATGTTGAACGACAAAATATAAGTAGATTAGAAAATCAAAAAT
CAACTGCCGAAAGACTGAATAAAGAATTGCATGATAAAATTGAAGAATTGGAAATGGAAGCTGGTAAAAAATATAAAGCT
ACAGTTGCTGCTCAACAAAGTAAAATCCAAAGTCTTGATTCTCAGCTGGACGCTAAAATTGAGGAAATAAACCAAGCTAA
TAGAAATAACAAGAAACTAGAAAGAAAGTTGAAAGAAATTATGGCTCTTATGGCTGATGAAACAAAGCGGGCTGACAATG
CAATCGAACAGATTGAAAAAGCCACTGGCAGATTTAAAAAAGCAGAGAGAAGTTTACAGATGGAACAAGAAAACAATTCT
GCTCTAAGTACACAAAATCGCCGATTACAACGAGATTTGGAAGAAGTAAATGAAACCAAAGAAGCACGGGACCAAGAAAT
TAAAATTCTTAAAACCAAAATCGACAGATTGGAAAAAAGAACACGCGCTGGGGGTCGACCTCTTAGAGGAGACGGTGGAT
CTGATGATGGGTTATCTGGAGATGGAGAATCCGTAACTGATTCTGCTTTAAAATCTGTTCAAGAAGACTAATTCAATAAG
TCCTAAAATGAACAATAAATTCTTGAATAATTAGTTAATACAAAAATTTTTGAATAATTTATATTTGGACTAACTTTTAA
TTCCTATTTGGTTTGCTCAATATATAATTGTTTTGATATATTTAATATTTCTATTTTATCAATTTGTGTACTTGGGG
>SMED30002016 ID=SMED30002016|Name=POU domain protein|organism=Schmidtea mediterranea sexual|type=transcript|length=1339bp
GTTGAATTTTTCAAAAACCAAATTTACTTATTTTGAATTGCCATAATGGAAAATGGAGTTGCAACAAGCTGTTCACTTCT
AGCTAATGGTGGATATCATAACTCTGCCTCCGTCGGTTGGAGGTTTCCGCAGAACAATTCAGCCAGCGATTCCGTTGAGA
ACGAATTTATATCAAATGCAAATCTCAAAATTACCTCTTCGAATATTCCAAACTTTTTTTACCAATGTCCGGAAAACAGT
CAATCAACGAGCAATTCTAACGGATTTTATTCAGTGGATACGAATTGCTCGGCTGGATTCCAAGACATGAATACTTACTA
TTCAAGGGTTGCAGCAGGCTTCAATCAATCCTTCGACTTGATACCTCAACACCAGTCTTTTGTGAGGCTCAATTCAACTG
GGCTATCGGAAACTGACCAAATGACTTATATAAATCCAAATCTTCATAGTTCATTTCATCGAAATATTTCATATCCGAAT
TACGGTAGTAACTGTCCTCCTAATGTTAACATAGGAGGAGTTTCCATGGTTCAAGGGTCTTCGTCATCCTTTTTAGGCTC
CAATTATTCAGGCATGCAACAACCAATTCAAGCCGTGAATCAAGGTGACATTGAATATGAAACCCGAGAATTAGAAGCTT
TTGCTGAATGGTTTAAACAAAGAAGAATAAAACTGAGAGTGACTCAGGCGGATGTTGGCCGAGCTCTCGGGAATTTAAAA
TTAGCCGGTGTTGGATCATTGAGTCAATCAACAATTTGTCGGTTTGAATCACTAACTCTCAGTCATAACAATATGATTGC
TCTTAAGCCAGTTTTGCAAACCTGGCTCGAAGACGCTGAAACCGAAGCTAAGGAAAGATTAAATGGCTCTGATGCTTTCT
CTGACATAGATGAAAGAAAACGCAAACGAACTTCAATCACAGATTCAGAAAAACGATCACTGGAAGCATTTTTTACGATT
CAACCACGCCCATCAAGCGAGAAAATTATCCAAATTGCGGAGAAATTGAATTTGAAGAAAAACGTTGTTCGAGTATGGTT
TTGTAATCAGCGCCAAAAACAGAAACGAATGAAGTTTTCCACTCTCGGTTTTCAGCACAACTCGTCAAACATGTGAAATC
CTATTTCTGTACAGGACAAATACCAAGCATTGACCCCTCTGTCTTATTACAAAAAGTGGTATGATTAAAAGTAGAATAAA
AATGATTTTTATTATTTCTTGACCGTAGCTTTTGTTTAAATTGTTTGACTTCATTTTTACGAAACTGTTATTTTTCCCAC
TGGACTGAGTCTTAGGTTCAAATGTGAGATTTGAGGTTGAGCTTTTGCTGAGAATGACA
>SMED30001362 ID=SMED30001362|Name=SMEDWI-3|organism=Schmidtea mediterranea sexual|type=transcript|length=3093bp
AATTAAGAGACAGCCATTTATGTTTATTTTTAAAATAAATAGTATATATTTAAGTTTCGAATTTTCAGCTACAATTTAAT
AGTACATAATTATACGATGTCAGGAAGTAGTGGAATAGGTAGAGGCCGCAGTCGTGGGCTGTTGATGCAAAAGTTTCTGA
ATAAAGATGTTCTTGTTCCTTCTGTTGAATCTTTAGAAGACAAAGCTCTTAATAAGCTAGGAATTCCACTTCCTGGGTCT
ACCGTAGAAAAAAATACAGAGTCAAGTTCTATATCAAGTGGAGATTCAAGAAGTAATCCCAGTGGAGATTCAAGAAATAA
CATAAAACTACAAGATAGTGATATTGAGAATCGCAATATTACCATTGTTACGCGACCATTATCCTGCATAGGCAGGGGTC
GGGGTTTAAGCAATCCCTCAAGTTTAACTACGTCATCGGGTAAATCGGATAAAATTACTGAAAACGAAGAACCTGGTCAA
ATTAAAAAGTTTGTAGGTCGCGGTAGAGGATTGTTGAATTCTCAGAAAGAATGTTCAAACTCTACTCCATCTGAAGTTTC
AAATGAATTGAAACAAATGAAAATTTCAAATGATGATAAAATGACGGTTTCTTCAGAAGCAAAGTCACAATTTGAGAACA
TTGAAAAACCTATTAGTAAATTTCGTCGACGCGAATATCCAACTCAAATAAAAGAACCATGTAATACAAGAAATGATTCA
TCGCCATCTTTAACTTTAAGTGCTAATTACGTTAAAGTTAGGACTACACAACCCCATATATATCAATACCATGTTTCCTT
TGCACCTCCGATAGATTCAAGGTTGATGAGAATTAAAATAGTTCAAGGGTTATCAGAATCAGATTTAGGGGTTGTCAAAG
AAGCAAGAGCTTTTGACGGTATGAATTTATACATTCCTCAACTTTTAAAAAATAAAGAGACAATAATTAAAGTAAACAAA
CCGACTGATAAGACTGTCGTGGATGTTAAAGTAGTTTTTACTAACAATGTTAATTTTAGTGAATGTCCTATGGTTTATAA
TGTTCTTTTTAAAAGAATTGAAAATTCCCTCAGAATGGTTAAAATTGGTAGGGATTATTTCTACCCTGAAAAAAAGATAG
TACTTGACCGTAGAAGGATGGAAATATGGCCGGGATATGTAACAAGTATCCAAAATTTTGACGGTGGTTTACTGTTACAA
TGCGATGTGTCACACAAAGTTATTCGAAATGATAGTGTGTATGACATAATGATGGAAATTAATAAAACTGTTAACAATAA
AGGTCAAATGCAAACTGCTGCGATTAATCAACTATTGGGTCAAATTGTGTTAACTCCTCACAATAACCGAAATTATAGAA
TTACTGATATAGATTGGGCTAAAAATTGTTTAAGTGAATTCGATAAAGGAGGCGAAAAAATTAGCTACCGGGATTATTTT
AAGAACACGTATGGGCTACAAATTCGTGATCTAGAGCAGCCTATGATAGTTAGTAAATCTAATAGCAGATCTGGTAAAAA
CCGAGGTCCCAAAGGATCAAAAGAAGTGGATGGTGGATTGGTCTATTTAATTCCAGAATTGTGTATGCTAACTGGTTTGA
CAGATGACATGATTAAAGATTTTCGTTTGATGAGAGAATTACACGAGCATTGTCGAGTTACTCCCAAGAAAAGACACGAA
GCCTTACTGGAATTCGTGGATAACATATATAGCTGTGAGGAAGCTAAGAAACTTTTAGGGTATTGGGGTATAACGATTGA
AAAGGACACTGTCAACATAAATGCTTGTAAAATGAATCCAGAAATGATATATTTTGGAAATGAAGCTTCTGTTAGTGCTG
GGGAACAAGCTGAATTTAAACAAGCCTTGGCACATAATAAAGTTATAGGTGGTATTCGTATTGAAAATTGGATATTAATT
TCTCCAAAAAGTTTACTGACAAAAGCAAATGGTCTGTTACAGGCTTTAATGAGCAAATCTCCTAGAGTTGGAGTTATGTT
TGGAAAACCCAAAATAGTTGAAATGAACAATGATCGAACAGAAGAGTATTTAAAAGAATTAAAGAGAAATGTGGCTCCTG
GTGTGCAGTTAGTAGTTACAATTTTATCTGCTGTTAGAGAAGATCGATACAATGCAATAAAAAAATTTTGTTATGTGGAT
TGTCCTGTTCCAAGTCAAGTGGTATTAGCCCAAACATTGAAAGAAGGGCCTAAATTAAATAGTGTGGCAGTTAATATAGC
CCTTCAAATAAACGCAAAATTAGGTGGAGAGCTGTGGGCTGTCAAAATACCTATTAAGAAGTTTATGGTTGTTGGACTTG
ATGTTTGGCATGATACTAAAGGGAGAAGTAGATCAGTTGGAGCCGTAGTTGGTTCAACTAATGCGCTATGCACAAGGTGG
TTTTCGAAATCGCATTTGCAAGAACAAGATAAAGAAATTATGTACGTATTACAGTCGTGTATGTTAAGCCTTTTAAAGGC
TTATTTTGAAGAAAATAATTTTTTGCCTGAGACTATCTTTATGTATAGGGATGGTGTTAGTGATGGTCAGTTAGGATATG
TTCAAAAAACTGAAATTGAACAATTCTTTAAAGTTTTTGAATCGTTTAGTGCTGATTATAAACCTAATATGGTATATAAT
GTTGTTCAAAAGAGAATAAATACTAGGCTCTATGTAAGTGATCCGAAAAATAAAGGACAAATAAATAACCCCAATCCTGG
TACAATTGTCGACCATACTGTTACGAGGGCTAACCTTTATGATTTTTTTCTTGTTTCTCAATCGGTTAGGCAGGGAACTG
TAACTCCGACGCATTACGTTGTTTTATGTGACAATTCTAAATACACTCCGCATCAGGTTCAGTTGATGGCTTATAAAACA
TGTCATATATATTACAATTGGCCGGGAACGGTTCGAGTACCAGCACCTTGTATGTATGCTCATAAATTGGCATATATGGT
TGGTCAGAATTTGAAAGCTGAACCTAGTAATCTTTTATGTGACAGACTTTTTTATTTGTAACATATGTGTTTCTATTTGT
ATTATTGTTAAACTGCATTTCGTAAAATAAATTAATTTCGTTATAAAAAAAAA
>SMED30002072 ID=SMED30002072|Name=Intermediate filament protein|organism=Schmidtea mediterranea sexual|type=transcript|length=2324bp
CATAAATGGTGCACTTTTGAGAAATTTGCGTTGGAGTAAATAGAAAGCAGTCTTAACAAAAGGATAACTAGGTCAGAACA
ATAGCTGCGTTGAAAAATCTATTATAAATATATCCAACTTTCAACTTGTTTCTCTAAAGTTATTTTTGTAATAATGGAAT
CGTCTAAAAGAACTGTCATTACGACTTCTAAACAAGAAGGGCCGATTAGTGGCTCAAGTTCGGTTACTAGATCTGGAGTC
TCAATGACATCAGGTTCTGGTTCTACTTACGGATCGGGATCTGGAGGAGAATTGGGCTCAGGTTCAGGATCCGGCTCTGG
AAGTCGATATCAAAGTTCGTCTCGTGTTACTACAACAACTAGATCTGACTATCCACAATCTGGAAATGAAAGAGAAGACT
ATAATGAATCAGGAAGTCAACAATTCAATGAAGATGGATCAGAGATTTCTGGTCGAATCGTTTCAACTACCAGATCCATG
TATAAGCCAATAAGTCAATCAAGGAATAATTTCAAAGACTTGCAAAGAAATGCTGCATCTTTAAGTTATAGTATGCAGTC
TGGAAACCAACAAAGTGGTTCAGTTATGCCACTCATCAGAGGTGATAGTGTAGCCTTACAGACTGGACTGAATTCGATAA
AGAAAGGAAGAGCTCAAGAAAAAAAAGACTTACAAGAGCTGAATGAAAGATTTGCGAGTTATATAGAAAAAGCACGATTT
TTGGAAGAGCAAAATAAGCAATTAGTCGGTGAAATTGAAGAACTCAAAGGGGGAACTATTCAAGATACATCAAAAGTTAA
AGAGATCTATGAAGTTGAAATCAATCAACTCCGATCTTTACTAAACGAATCAGAGAAAAGTAGAGCACCACTGGAAATTC
AAGTGAATTCCTTGGAATACGAAATAGGCGATTTGCAAAAAAGACTGCTGCTATTAGAGGATATAGGTACACAGTATAGA
GCAACGATTGATAAACAGCGTCGAGAACTCGAGACCTTCGAAGGGGAAATGGTTGAATTGAAAAGGTTTAAAGACACTCG
AAGTATCGATGCTACAAAAGAAAAGGATGAAACAAGCAACTTAAGAGATGAAATAAAAAAATTAAGACAAAACTATGAAA
AAGAAGCATTAGCGCATATCACCCTTGAACATGACTTGCAGAGTTTGAGAGAAACTATGGAGTTCGAGAAACAAATCCAT
GCTACAGAAGTAAAAGAGCTGGCAACCGTTGCATATCAAAACCCAACGAAAGAAAACATGGTTTACTGGCAAAGTGAAAT
GGGACGTGCTTTACGGGACATTCAAACTGAATATGAAAACAAGCTTACAGGAATCAAACAAGAATTGGAGGCTTCTTATG
GTTTACAAATGCAGGAAATACGAACAAGTAAAACCAGAGACAATGTGGAAACGACCCATTACAAAGAAGAGAACAGCCGA
ATAAAATCTCAAATCCAAAGCATGAGATCGAGAATCGCTGATCTTGAGATGCAGAATGCTGAATACGAACACAACTTGAA
TAATATTCGTGCTAATTACGAAGAAACTCAAAGAAGTCTTGAATTGCAGTTGTTGTATTTGCAAGAGCAATTGACAGGAA
CTCAACAAGAATTGACAGTCGCTTTGACCGAACTCCAAAGCTTATTGGACGCGAAGTTGGGACTGGAATTAGAAATCGCA
GCCTATGCAAAACTATTAGAATTCGAAGAGAGTCGTGTCAATTTCTCTGGTGATGGCAACGCTACTTTATCTCAATTTAT
GAGTTTCACATCAAGTAGCGGAGCTGGAGGTGCTGGAGGGGCTGGAGGTGCTGGTGGTGCTGGAGGAGGAGGAGGAGGAA
GCAGTGGGTTCGGATCAGGAAGTAGAATTGGAGGGTCAAGTGGTTTCAGCGGAATGACAGAAAGAGAAACAACTGACTAT
CAGTCAAAACGAATGAAATTTAGACGAACGACAAAGGGAAACATTTCCATTGCAGAGACTGACCTTATAGGTTCAACAAT
TATTCTGGAAAATATTGGGACCAAAGTGGAATCACTCGAAGGTATGACTATTAAAAGAGTAGTCGACGGCAGCGAAAATA
AAATCTCATTTACCATTTCCGGAAAAGTTAACTTAAATGCTGGTGAAAAGATTTCTCTTTGTGCCAAAGGGAAAATAACC
GGCAAAGGACGAGAAATCGAAATTGATGATGCTTCATGGGGGGTTGGTAACAATGTCGTAACAACACTTTGTGATTCAGA
AGGAGCGGAAAAGGCAGCTCACTATCAAGAGGTTTCGGCCTAGAAACCTTGCTAAATATTAAACTGAATTTGTATGAACA
TAAC
>SMED30010115 ID=SMED30010115|Name=ribonucleoside diphosphate reductase subunit M2-2|organism=Schmidtea mediterranea sexual|type=transcript|length=1127bp
GTTTTTAATGAACTGAATCAGGTTTAATGCAAAATAGTCATTTTAATTTCTAGATAAATTAACAGTAATTCTTTCGAGAT
TAGATATTTATTATGGAGATTTTATTAACAGAAGATAAAAGTCGAATAGGATTTCTTAATATACAATATGAAGATATATG
GGAAATGTTTAATAAAGCTCGATCTTCATTTTGGACTGTAAATGAAGTTAATTTATCAAAAGATTTTGCTCACTGGAAAA
AAATGAATGACGATGAACGGTATTTCATTACTCACGTTCTTGCATTTTTTGCTGCCAGTGATGGAATTGTCAATGAAAAT
CTCGTCGAAAGATTCATTCAAGAAGTGAAAATTCCCGAAGCTGAGGCATTTTACAGTTTTCAAATTGCAATTGAAAACAT
TCATTCAGAAATGTATTCTCTATTAATCGATTCTTATGTTACTGATACAGAAACAAAGAATAAATTATTTAATGCCATCG
AAACTATGCCATGCGTTCAGAAAAAAGCTCAATGGGCATTGCGTTGGATTAGCGATAAAAAATCCACTTTTGGAACTAGA
GTTGTTGCATTTGCTGCTGTCGAGGGAATATTTTTCAGTGGTAGTTTTGCATCGATATTTTGGCTTAAAAAGCGTGGATT
AATGCCCGGATTATGTTTCAGCAACGAATTAATTAGCCGAGATGAAGGACTTCATTGCGATTTTGCTTGTTTGATGTTTA
AATATTTAAGTAGCAAACCGTCACCTGAACAAGTTTTGGATTTGATCAAAGAAGCTGTCGGAATCGAGAAGGAATTTCTC
ACCGACGCATTGCCGGTTTCATTGATTGGCATGAACGCTGAATCGATGTGTGAGTATATTGAATTTGTGGCCGACCGGCT
GCTGGTTGAATTGGGCTGTGACAAGTTCTGGAACACAGAGAACCCGTTGGATTTCATGGATAATATTTGCATCGATGGGA
AAACTAACTTCTTTGAGAAACGAGTCGGTGAATATTCCATGTCGAGCGTTGAGGACAAACACAAATTTGATGATGTCAAT
TTCGATATTGATTTTTGAGGGATTCCGTTTTTTGGCGCCAGTGTGCATTTAATGTGTTTTGGTTGTAATTGGCTATTTTT
GTTTGTT
>SMED30024865 ID=SMED30024865|Name=Intermediate filament protein|organism=Schmidtea mediterranea sexual|type=transcript|length=2214bp
GATCTTATAAATACAACATTAATTGTATAGTGAACTATGTTAGCCATTGAAATGGAGACAAGTAAAAAAATCGCAGTCAA
AACTACGAAACAAGAAGTAAGTGGAAGTGGAAGCTCTGGTGGACTAAATGAACAACAAGCTTTTGGAAGTGGTAGCTTTC
GAACCGGGTCTGGAGGTGTACAGATAAGTGCTAGTGGAGCAATTGGATCTAGCAGTTCAGGCGGAGTTAGTGGCACTGGC
GGAGTTGGTAGCATTGGTGGAGGCAGTGGTGCTGGTGGTGAAGGAACAGGAGCTAGCACAGTTATCACTACAAGCAGATC
TTATTATAAGCCTATCAGTCAACCTAGAAATAATTTAAAAGACTTACAAAAAACTTCTGGTATGAGTTACAGTGCACAAT
CAAACTCCCAGCAAGGTTTAAGTAATATTCCATTGATTAGGGGAGACAGTGTTGCTCTACAATCTGGAGTAACTTCAATG
AAAAAAGGACGAGCTCAAGAGAAAAGAGATCTTCAGGATCTGAATGAAAGATTCGCTAGTTACATTGAAAAAGCTCGATT
TCTAGAGGCGCAAAATAAACAGCTAGCCAATGAGATTGAAAAGTTAAAAGGTGGAACCGTCCAGGACAACTCAAAGATCA
AAGAAATCTATGAAGTCGAAATCAATCAACTTCGAGCTTTGGTAAATGAATCGGAAAAAAGCAAAGCTCCTTTAGAAGCT
CAAGTTGATTCTCTTGAATACGAAATTGATGATTTGAGAAAACGATTAATGATGCTGGAAGACATCAACTCTCAGTACAA
ATCCACCATCGATAAGCAACGCAAAGACATTTCTGGATTTGAAGGGGAATTGGTGGAGCTAAGAAGATTCAAAGGCACCC
AAGGTGCTGATGTAAATAAAGAAAAGGAAGAAGTAAAAACACTAAGAGACGATTTGAAAAAATTGAGACAAGATTATGAG
AAAGAAGCTCTAGCTCATCTCACCCTTGAACATGAATTACAAAGCTTGAAAGAAGCATTAGAATTCGAGAAACAAATACA
TGACACAGAAGTAAAAGAGCTGGCAACCGTTGCATATCGAGACCCAACGAAAGAAAATATGGTTTACTGGCAAAGCGAAA
TGGGACGCGCGTTACGAGACATTCAAACTGAATATGAAAACAAGCTTACAGGAATCAAACAAGAATTGGAGGCTTCTTAT
GGTTTACAAATGCAGGAAATACGAACAAGTAAAACCAGAGACAATGTGGAAACGACCCATTACAAAGAAGAGAACAGCCG
AATAAAATCTCAAATCCAAAGCATGAGATCGAGAATCGCTGATCTTGAGATGCAGAATGCTGAATACGAACACAACTTGA
ATAATATTCGTGCTAATTACGAAGAAACTCAAAGAAGTCTTGAATTGCAGTTGTTGTATTTGCAAGAGCAATTGACAGGA
ACTCAACAAGAATTGACAGTCGCTTTGACCGAACTCCAAAGCTTATTGGACGCGAAGTTGGGACTGGAATTAGAAATCGC
AGCCTATGCAAAACTATTAGAATTCGAAGAGAGTCGTGTCAATTTCTCTGGTGATGGCAACGCTACTTTATCTCAATTTA
TGAGTTTCACATCAAGTAGCGGAGCTGGAGGTGCTGGAGGGGCTGGAGGTGCTGGTGGTGCTGGAGGAGGAGGAGGAGGA
AGCAGTGGGTTCGGATCAGGAAGTAGAATTGGAGGGTCAAGTGGTTTCAGCGGAATGACAGAAAGAGAAACAACTGACTA
TCAGTCAAAACGAATGAAATTTAGACGAACGACAAAGGGAAACATTTCCATTGCAGAGACTGACCTTATAGGTTCAACAA
TTATTCTGGAAAATATTGGGACCAAAGTGGAATCACTCGAAGGTATGACTATTAAAAGAGTAGTCGACGGCAGCGAAAAT
AAAATCTCATTTACCATTTCCGGAAAAGTTAACTTAAATGCTGGTGAAAAGATTTCTCTTTGTGCCAAAGGGAAAATAAC
CGGCAAAGGACGAGAAATCGAAATTGATGATGCTTCATGGGGGGTTGGTAACAATGTCGTAACAACACTTTGTGATTCAG
AAGGAGCGGAAAAGGCAGCTCACTATCAAGAGGTTTCGGCCTAGAAACCTTGCTAAATATTAAACTGAATTTGTATGAAC
ATAACAAAATTTTTAGTAATTTAATAAAAGTTTATTATTTTATAAATCCAAAAA
>SMED30010910 ID=SMED30010910|Name=POU domain protein|organism=Schmidtea mediterranea sexual|type=transcript|length=3203bp
TAGGGCTAAATGTAACGCTACTTATCGGTATTGCCACAATTATATTAAGGCAGTGGCACAAACTTATTTTCTCTACCAGG
CCTGCTGAATCGACAAACAAACATATTCATCGTCATTAAATGGGCAGTAGTAGTAGAAACACAGTGGTAAAGACATGGCA
AACACAGCAGCATCAGTGATAGCTGGAAGAGATAGCTACAGCATCAGCAGCAGTAACTCTCGCAGCTAATAATGAAGATA
ATGAGCTAGACTTAATTATCTTGGCTAATAATTTCAAATAAAATATCGCAACTAATTTTGTCATTCGTATTCTGTGAGGA
TAAATCACAAATTTGTTTCGATTGTTGGAAATTTAAATTTTTTTCCCTCGTATTTTCCAAATACTCCAGAGTTTGAAGAG
TATAATGAATTTGTTACCGAATCCGTTACTCACAAATGTGCTGGATATGACTAAACTTGACCAACAAAACCAAATTATCT
CGTGTGGCAATTCGACAAATTTCACCAACAAAAACACAATTGACAGCTCTTCAAATGATACCACACAAGTCAGTTCAACA
GCCAACAACATCAGTTCAATTGAACATTTCCTTTCATCCCGCAATTTAACAAATCAGCTGATAAATAATTCCGGACTCCC
ATTAGGGGCACAGATATTCACCATTCCGGCATCATTGGCGACATTCCCGAATTCGGATTTGATTCCAGTATTCATCAACA
GTTTAGGACAAATGGTTCTGGCAAATACTGGGAATATTCTGGCGAACAGTTTGGCAAGTTTGGCCACCACACTCACGTCA
ATCCAGCAGCTGTCCGTGAACCGTGAATCGAATGTCATCAACACTAATCAAACTACTAACCCAAATGCAGCTATCATAGA
GCACATGATGCAGCAATACCAACAGCTTCAGCTGCTGAAGAACTCAATGATCCATCAATCATGGCCGAATGAGTTCCAGC
AGACATCCTCAAACAACCAAACCAATTCATTTATTGCAAATAATATACCAGTAGCAACTACAAATTCTAGTACAATTAAA
CCCATAGATAAAGTTAATGGCAGTACGCAATTCCAAATTCCGTTGAATTGCAATCCTGCTCCTCTTCAAAATGAAAAATT
TACTACCAATATCATCTCCATAAATGATCGTGCTATAGATAACAAATCCCCAAGCGTCAACTCTTCGGTATCCTCTAGAT
CAGTAGGAAACAAAATTTGTCAAATGATTGACGTCAGTCATGATTACATTTTAAGTCAAGAAGATCATCTGGTGAAAGAA
AAAGCCGCAGAATTACTCCGACATTCACACAACCAAAAAAACGCTATCAAACATGCCCAATGTTCTGATAAATCAGAACC
AACAGCACATAATGTCACAGTGGATGGTGTTGATCTCGGTGAAATCAAAGAATTTGCCCGTAATTTCAAAATTAGACGTC
TCAGCATGGGGTTAACTCAAACACAAGTCGGTCAAGCGCTGACTGCAGCAGAAGGTCCGGCATACAGTCAAAGTGCAATC
TGCAGATTCGAAAAGTTGGATATCACACCGAAATCGGCTCAGAAAATTAAACCAATTCTAGAGAAATGGATCAAAGAATC
CGAAGAAAAATATCTCAGTGGATCGAACAGTGTTGATGATGGATCCGGATCTGACATCAATAAAAAACGGAAACGTCGAA
CGAGTTTTACACCACAAGCACTGGAAATTCTGAATATTCATTTTGAGAGGAAATCTCATCCGACTGGATCGGAAATGCGA
GCATTGGCTAACGACCTCAATTATGACAGAGAGGTCATTAGAGTTTGGTTTTGCAACAAAAGACAAGCGTTAAAGAATTC
TGCCAAGCGAATTAAACAATGTACACCAAACAGCGAATCAAACAACCAGCACATATCTCCCCAAGTCATGAACAGCAACA
GCAACAGCAACAACACCACCATCACCAACCGGGCAGTCACCTAGGCCAAATGCATCCATCCATGATGGACATGTATCAGC
CACACGCGTTCTCAAGACCCATTACTGCTGAATACCAACAGAGTCTCAATCGGCAACAATATCGACGCAGCGACAGCGAT
TGTGGAACTCATGAGCCTGGATATTCGTACCTGCCGACTCCTCATGGGCCAGAAGTGGTGTCATCTCGTCAACATACTCT
CAATAGTCTTCCAGAACAACCGATGGACAATAACAATCATGATCAGCATGATTCTCCAGTACCGATTATTGCCAACAGTA
GCAACAGTAACATCAACAATAACAACAGCAGTTCTGGTCGACATCATCATCGTCGCAAAACCAGATCTCGACAAACCAAT
AATCAAGAGCCGACATCACTAGAACGGAGCCAAGTAATGAAAAATCACAACGACATACCACCAATGTTAGCGCCACGGAC
AAAATTGCCGTATTCCGAGCATCAGGGAATTCATCGTCTATCCGCTCATGAAATGAATACCATTGACAGTCGTGACAATG
CTGTCAATGATTGCAATTCTGAATCCAATACCGCTGGCGCTGTCAGCGATAATTTCGACCCTCCTCTTCATACTCCGGCC
ATTACGATACCAATGAGTCGAAGTACCGCATGGATTGCAACATCGATATGAATGAAAATGAAGCAACTCCCAGTAATTCT
CTGTTGATTGAGAACCTTCCCAGAGTTAGAGATGTTTCCGAGTATGGAATACCTATAAACAATAACAATCACAACATGAA
ACAAGCTCCCGTGCCTCCTCTTAAATCGAATAGTCGACATAATCCTGTGCCTAACAACAGTTACGGTCTCGAGTGCAATC
CTGATTATTATTCATCTGCCACATCTAGTGACAGATCCGGTCCCAGTCCTCCCCCTAAATATGCGCCAGGAGTAGTTCTT
CCTGCCCGTGCTGATGAAATATATATGCGCGATCAAAGAGAAGGAATGTACTATCACGGAGATACTCATTGTTGATAATT
GTATTTCACAATTTCGTTTCATCGGGTGTTCTCGATTGCTGATACTCTATCTCATACCTTTCGATGAATTTCTGATCCCA
ACTGACAGTTGTACAATTCTTATCATCATCATCATCTTCCTCATCTTTTCATATTTTATCTAGTCCTACTTGCAATCTTC
TCCGTGTGCTCTCCACAGTGCGATTATTATTATAATAATAAATTATTGCTATTGATTATTATAATTATTGTTATTGATAA
TAA
>SMED30030594 ID=SMED30030594|Name=Opsin|organism=Schmidtea mediterranea sexual|type=transcript|length=1880bp
AATAAAATGAAATTTATAATGCCATTGGTATTTACACTGTGGTTAATAACGATAATAATCAGAATTTTTAATGATAAATA
TAGTAAACGATAACTAGTAACGGTAACCCTAATAATGAATGCTATATTACATATAGTAGTGTTACCAAAATTTTGGAATT
TAAAATAATAAGTTTATTATGAGTGACCATGATAATAAAAAAGATTTTTAAATAAAATGATAATATTACTATATAGTCGG
TTATAGTTCCGATTAGTCTACTATCGTAGACGTAATAAAAAGAGTTTAGTAACCTAATTTTCAATGTTTTTTGAAAGTTG
CTCGTCTCTATGGCAACTTATTATCGTAAGGAAAATTTTCTACTTCACTATGATATTTTCGGAGTGAATTTATTGATTTA
CATTCAAGGACTTTCACTATTGCATTTACTGAAGCGATCGTTTCATTTGGAACTCATATTTTCGCCCCTAATCGTTCTGG
AATTTATTATTTATGATCCTTTATCTGACCTTAAAAATTTTCATATTCATGAGTGTCCAATCTTTTGCCAATTTGGGAAA
TAAGCTCCTAGAAAATGCTACCTTCAATAATGAATCCCTTACAAAGTCAGTATGGCATTGGGATCCTGAATTTGAATCAA
TTGTCCATCCGTACTGGCGGACTTTTGATATGGTGCCAGAAGTTTATCATTATTTAGTTGGAGTGTATATTTCTATTGTT
GGAATATCGGGAGTTCTGGGAAATCTTCTTGTTCTTTACATATTTGCAAGCGCCAAAAGTTTAAGAACTCCTCCAAATAT
GTTTATAATGAGTTTAGCAATTGGAGATTTGACATTTTCTGCTGTGAATGGCTTCCCCTTACTTACTATTTCAAGTTTTA
ACACTCGATGGGCTTGGGGAAAATTAACGTGTGAGATTTATGGTTTCATCGGTGGTCTTTTTGGGTTTATATCCATCAAC
ACAATGGCACTAATTTCTCTGGATAGGTATTTTGTTATTGCCCAACCATTTCAAACAATGAAATCCCTGACAATCAAAAG
AGCAATAATCATGTTGGTTTTCGTATGGCTTTATTCATTAATTTGGTCAACACCGCCATTTTTTGGATATGGAAATTATG
TGCCGGAAGGATTTCAAACGTCTTGTACATTCGACTATTTGACCCAATCAAAAGGTAACATAATATTCAATATTGGGATG
TACATAGGAAATTTCATAATTCCCGTTGGAATAATTATATTTTGCTATTATCAAATTGTCAAAGCAGTGCGAGTACATGA
ATTAGAAATGTTAAAAATGGCTCAAAAGATGAATGCATCTCATCCAACTTCCATGAAAACGGGTGCAAAAAAGGCTGATG
TTCAAGCTGCAAAGATTTCTGTCATAATTGTCTTTTTATATATGTTATCATGGACACCATATGCAATAATTGCCCTTATG
GCTCTCACAGGGCGTAGAGATCATCTGAATCCATACACTGCAGAATTGCCGGTACTCTTTGCTAAGACCTCAGCTATGTA
CAACCCATTTATATACGCAATAAATCATCCAAAATTTCGAATTCAGCTGGAAAAGAAATTTCCATGTTTGATTTGTTGCT
GTCCACCAAAACCAAAAGAAAGGGACACTACAAGTGCAGCATCAGCAACGAAACCGAGTAAATATCAGAGTGAAAGAGAA
AGTTCAATGTCTCAACTGGATCCTGAAGACGATAAACAAGTCGTAGAAACCACCAAGAATCAAACCACAAGTGGACATAC
AAATGAAGCTTACGAGAGTGAACAGAAGTAGTTGATGAAACTAAATTATTCTATTTTTAATGACACTAGATCTTTATAAT
AATTTTATTTAAGGGAATGAGCACTGAATTTTGCGCAAAT
>SMED30018592 ID=SMED30018592|Name=ovo|organism=Schmidtea mediterranea sexual|type=transcript|length=1211bp
GTCATATTGAACACTGAGACTAGCTATCAATAGCTGACAATTAGACGTATAAAATGTTAGGGTACGCCTTTTTGGCCATT
ATTTCACACCAAAATTTCCATGAGTATTTTTCTAATAAGCACTCTACTCGCAAACGATGACGAAATGCCCACAGATTTGT
CATCGAAAACTGAAAAGATTGAAAACCATGAAAAATCTGAAAGGAATCTCTGCAAGTATTCCAGTCAGTTTCTTTCCTGT
TTTTATTCACAGGAAATGTTGACACTACTTATGAGAAAAAATTTCATCAATCAATATTCATTTTATTTAAAGATTTTCTC
GGGGTTTCCACAATTCGTCGATAACTTCAACTTAGTAAATCCAACAGATTTCACCGTCAATTCAAATTCAACAACAATTT
TGCCCTCATTCAACTTCCATAAAACGTCATTGAATATCGAAATGAACACCTCACTGGCCAATTCTTCAAGCACTGGTAGG
TCGTCTCCAACTGAAACTGTGTCTGAAAATCTTTGGTTAGAAAAAGTAAAGGTGGAAAAATATTTCCATTTAATTGAAAA
TCTATCACAATCCCAAGAGAACATTGAAGACATTGTCAAAACGATCAAAGTCAAACGAAGAAATTTTGCTGAATATATTA
AGAAACTTTACCTAAGTGATCCAGCCAAATTTAAGTTGACAAATGGTGGAGATGGAATAGTAAATCCATTTAGAAAACAA
TTCAAAGAAGAACGAGATAAATCTATGGACAAATTTTGCATCGAAGTTAATCACAATTATCAATGTAAGATATGTAACAA
GGTGTTTCCTTCCAAAAAGCACATGCAACGTCATATACGTTCACACGGTGTAAATTTTGATTGGTTGTGTAAATATTGTT
TTAAACCGTTTATTGATTCCTACGATTTAAAACGACATACGAGAGTCCATACAGGAGTTCAACCATACAAATGTCAAAGT
TGCACAAGAAAATTCAGTCAACGATGCTCATTGGAAAGTCATCAGGTGAAAATCCATGGAGTAGCACTGAATTATCTATA
CAAAGAAAGAAGGAACAAGTTGTATCCGTGTGAAATCTGCAGTTATAGTACGTCTTGTAAAGCAACTTGGTTGAGTCACA
TCATTCAGGAGCACCCGAATTCACTTTATGCAAAAGAAGAAGCACTCAAAGGAAAATACAACTGACTTTGAAAAATATTG
ATTTTTCAAGC
>SMED30025388 ID=SMED30025388|Name=Myosin heavy chain|organism=Schmidtea mediterranea sexual|type=transcript|length=6089bp
TTTTTTTTCGAGTTTGTGCTAGAAATTAATTTCAGTTTGAAATTTTTTGCCTATTTCGAGTTCTTTGAAAACAATCTGAA
AACATGAACCCATCTGATCCCGATTTTCAATATCTCGGTGTCGACCGAAAAGCTTTAATGAAGCAGTATGGGGATTCCTT
TGACAGCAAGAAAAACTGTTGGATTCCTGACGAGAAAGAAGCCTTCATTGCGGCGACAATTGAAGATGCGTCTGGAGATG
TCTATACGGTCAAAACCGATAAAAGTGAAACCAAGACTGTTAAAAAGGACGAAATCGAACAAATGAATCCACCGAAATTT
TTGATGATTGAAGACATGGCAAATTTAACCTTTCTAAACGATGCCAGCGTTCTTGATAATTTAAGACAAAGATATTACAA
GAATCTGATTTATACCTACTCTGGACTGTTTTGTGTGACAATAAATCCATACAAAAGATTTCCAATTTACACCCCGCAGG
TGATTTCAAAATACAAAGGTAAACGACGAACTGAAATGCCTCCACATATTTTCTCGATATCCGACAATGCTTACACCAAC
ATGTTGGCGGACCGAGATAATCAATCTGTCTTAATTACTGGTGAAAGTGGAGCTGGGAAAACTGAAAATACAAAAAAAGT
TATCACTTATTTTGCACACGTTGCAGCTGCTACTAAAAAAGAAGAGGATGATACTGGAACTGGTCAAAAGAGAAAGGGAA
CATTGGAAGACCAAATCGTGCAAGCAAATCCTGTTTTAGAAGCATACGGTAACGCAAAAACTGTGCGAAACAACAATTCA
TCCAGATTTGGTAAATTCATTCGTATTCATTTCGGTACAAGTGGAAAAATAGCTGGAGCTGATATTGAATTTTACTTACT
CGAAAAATCGAGAGTAAATTCTCAACAGAAAGGTGAAAGAAACTATCACATTTTCTATCAAATTTTATCGAACGGAGGAA
AACCATTTCATGAAAAATTACTCATCTCTCCTGATCCAGCTTTGTATTCCTTCATTAATCAAGGTGAATTAACAATCGAT
GGAGTCGACGATGAAGAAGAAATGAAAATAACTGATGAAGCTTTCGACATTTTGGGATTTTCTGCCGAAGAAAAAATGAG
CTTGTTCAAGTGTACTTGCTCAATTTTAAACATGGGCGAAATGAAATTTAAACAAAGGCCAAGAGAAGAACAAGCCGAAG
CCGACGGTACTGCTGAAGCTGAGAAAGTGGCTTTTCTGTTGGGTGTCAATGCAAAAGATTTAATGCAATCGATTCTGAAA
CCGAAAGTTAAAGTTGGTAATGAATATGTCACAAAGGGGCAGAATAAAGATCAAGTTCTTTATTCGGTTGGTGCGCTAGC
AAAATCTCTTTACAACAGAATGTTTGCGTGGTTGGTTTTGAGAGTTAACAAAACCCTCGACACTAAAGTCAAGCGACAAT
TTTTCATCGGTGTTTTGGATATTGCCGGTTTCGAAATTTTCAATTTCAATGGATTTGAACAAATTTGTATTAATTACACC
AATGAAAGATTGCAGCAATTTTTTAACCATCACATGTTCGTGTTGGAACAGGAAGAGTACAAGAAGGAAGCGATCGATTG
GGAGTTCATTGATTTTGGAATGGATCTTCAAGCGTGCATTGAGCTCATTGAAAAGCCAATGGGTATTCTGTCAATCTTAG
AAGAAGAGTGCATGTTTCCGAAAGCATCTGATATGACATTTAAAGCAAAATTGTATGATAATCATCTCGGAAAAAGTCCC
AATTTTGGAAAGCCAAAACCTCCAAAACCCGGACAGGCTGAAGCTCACTTCGAATTGCATCACTATGCTGGCAGTGTGCC
ATACAATGTTACTGGTTGGCTGGAAAAGAATAAAGATCCTCTCAACGAAACTGTGATAAATCTACTTGCGGCAAGTAAAG
AGGCATTAGTTTCGAGTTTGTTTGCTCCGGCTGAAGATCCAAATAATACGGGTAAACGGAAAAAGGGTGGGGCCATGCAG
ACCATTTCTTCAACTCATAGGGAATCCTTGAATAAATTGATGAAAAATTTACAAAGTACCAGTCCACATTTTATCCGATG
CATTGTTCCAAACGAATTCAAACAACCTGGTGTAATCGACGCTCATTTGGTATTACATCAATTGCATTGTAACGGTGTCC
TTGAAGGCATCAGAATCTGTCGAAAGGGTTTCCCAAACAGAATGATTTATTCTGAATTTAAACAAAGATATTCTATCTTG
GCTCCAAATGCAATTCCACAGGGATTTGTTGAAGGCAAACAAGTAACGAGCAAAATTCTGGAAGCTGTTCAATTGGACAC
CAATCTTTACCGCCTGGGAAATACTAAGGTATTTTTCAAAGCTGGAACTTTGGCTGATTTAGAAGACATGCGTGATGAAA
AATTATCTTCACTGATTTCATTGTTTCAAGCTCAAATACGCGGATATTTGATGCGAAGACAATATAAAAAATTACAAGAT
CAGAGAGTTGCTCTATCTATCATTCAAAGAAATATTCGTAAATATCTTCTTCTGAGAACATGGCCATGGTGGAAATTGTA
TACCAAAGTTAAGCCACTTTTGAACATTGCTCGTCAAGAGGAAGAAATGAAAAAGGCCGCTGAAGAATTGACAAAATTAA
AGGAAGAATTCGAAAAAGTTGACAAATTCAAAAAAGAACTTGAAGAACAAAACGTCAAATTGTTGGAAGCTAAAAACGAT
TTGTTCTTACAATTACAAACCGAACAAGACAGTTTAGCCGATGCCGAGGAAAAGGTTTCAAAATTGGTTATGCAAAAAGC
CGATATGGAAGGAAGAATTAAAGAATTGGAAGATCACCTTCTAGAAGAAGAAGATAATGCTGCTGGACTGGAAGATATCA
AGAAGAAAATGCAGGGTGAGATTGAAGAGTTGAAAAAAGATGTTGTCGATTTGGAATCTTCATTACAAAAGGCTGAACAA
GAGAAAACCGCGAAAGATCAACAGATAAAATCGTTACAAGATCAAATGGCAAAACAAGAAGAAGAGATGAACAAATTGAA
AAAGGAGAAAAAGGCCGCCGATGAAGTGCAAAAGAAAACCGAAGAATCTCTACTAGCGGAAGAAGAGAAAGTCAAAAATC
TAAACAAAGCTAAAGCCAAATTGGAACAGACCATTGATGAGATGGAAGAAAACCTCTCGCGAGAACAGAAAATCCGAGCT
GACGTAGAAAAAGTTAAACGAAAAATCGAAACTGAACTGAAACAAACTCAAGAGACCGTAGACGATTTGGAAAGAGTAAA
ACGAGAATTAGAGGAACAATTGAAGCGCAAGGAAATGGAATTAGTCAATGCTTCATCGAAAATTGAAGACGAATCGGGAT
TAGTTGCCCAGCTTCAGAGGAAAATCAAAGAACTTCAAGCTCGAATTCAAGAATTGGAAGAAGATCTGGAAGCAGAAAGA
CAAGCCCGTGCAAAAGCAGAGAAAAGCCGACATCAACTCGAAGGCGAGTTGGAAGAATTGAGTGATCGACTGGAAGAACA
AGGAGGCGCAACCTCAGCTCAAATGGATCTCAATAAGAAGCGAGAGGCAGAATTAATGAAATTAAAGAGAGATTTAGAAG
AATCGAATATGCAACACGATCAGGTTATCGCACAAGCTCGAAAGAAGCAACAGGATGTTGTCAATGAATTCTCAGAACAG
CTCGATCAACTGCAGAAAGTCAAAGCTAAAGTTGAAAAAGAAAAGAATGAGATGAAAGAAGATTTAAATGACCTACAAGG
ACAGCTGGAATCATTAAATAAAGCCAAGGCTAACAGCGACAAACGAATCAAAGAAATCGAAGCTCAAAATTCCGAATTAC
AGGGCAAATTGGAAGAGGCAAATAGACATATCAATGATGCCAACAATACATCTGGAAAGAATCAACAAACCAATGCGGAA
CTCCAAGCACGTTTAGAAGAAGCCGAATCGCAAATCAATCAGCTGGGTAAAGTCAAGCAACAAATGCAGACTCAACTCGA
AGAAGCCAGGCAAATCGCTGAGGAAGAGTCGAGAGCAAAGGCTAAATTGAGTTCTGATGTTCGCAATCTCAATGCCGATT
TAGACAATCTCAGGGAGGCACTGGAAGAGGAAAATGAGAACAAAAGTGACATACAGAGACAGCTAGTTAAAGCTCAGGGA
GAAATGCAACAATTGAAGGCTAAATTCGAAGGCTCGGGATCCATTCGTTCAGAAGAGTTGGATGAAGCCAAGCGAAAGTT
TATGACACGAATCCATGAGCTGGAAGAGGAAGCTGAAACCGCCAAATCGAAAGCCGGACAACTAGAAAAAGTTAAAGCTC
GACTCCAAGGAGAAATCGAAGATATGCTGGTCGACGTTGACAGAGCGAATACACTCGCATCTCAGTTGGAAAAGAAGCAG
AAAACATTCGATAAAGTCGTGTCAGAATGGCAGCAGAAATATGCTGAATCTCAGGCGGAATTGGAAGCATCCCAGAGAGA
AAGCAGGACGATTTCAGCTGAGGTGTTTCGAATGAAAGCACAAATCGAAGAAAGTCAAGAGCAGTTGGAAAGCGTTAAAC
GGGAAAACAAAAATCTAGCAGATGAAATCCACGATTTGACAGAGCAAATCGGAGAAGGAGGTCGCAGTGTTCACGAAATC
GACAAAGCAAGGAAACGATTGGAAATGGAAAAGGAAGAATTGCAACATGCGTTAGAAGAAGCCGAACAAGCTTTGGAACA
GGAAGAAGCCAAGTCTCAACGATCACAACTGGAAATGAGCCAGATCAGACAAGAAATCGACAGGCGATTGGCTGAGAAGG
AAGAGGAATTCGAGGCCACTCGAGTCAATCATCAAAGAGCAATGGAGTCAATGGAAGCCTCACTTGAAGCCGAATCCAGA
GGCAGGGCTGAAGCAACGAAGATGAAAAAGAAACTGGAGCACGATATCGGTGAACTGGAAGTTGCAGTGGATACCGCCAA
CAGAATCCGATCGGAGCAAGAAAAGAACGCAAAGAAATTCCAGCAGCAAGTTCAAGAATTGCTATCAATGGTGGAGGATG
AGAAACACCAGAAGGATCAGATACGAGAGCAAACGACGATGACAGAAAGGAAATTGACAATGGTCCTGGCAGAGCTTGAG
GAGGTTCGTGGCTCATTGGAAAATTCCGAGAGAAACAGGAAAAATACCGAGAATGAGAAAATGGAAATAACTGATCGATT
GAATGAGTTGAGTGTCCAGAGCTCCGGTTTCATGGCTACCAAGCGGAAATTGGAAGCTGATTTGGCTGCTATGCAGTCCG
ATTTAGAGGAGGCCGCTACTGAAGCCCGACAAGCCAATGAACAAGCCAAGAAGGCTGTTGCTGACAGTGCGAGATTGTTT
GATGAAATCCGGCAAGAGCAGGAACACGCACAGCAAATCGATAAGATCAAGAAACAACTCGAGGCCCAAAATAAAGAATT
GCAAGCCAAATTAGATGAATCGGAGAATAATGCTTTGAAAGGTGGCAAAAAAGCCTTGAGCAAACTAGAGCAACGCGTTC
GTGAACTTGAAGCCGAATTAGACGGAGAACAAAAAAGACACGTTGAAAGTCAAAAGAACTCCAGAAAAGTTGAAAGACGG
CTTAAGGAGCTTGGATTACAAACCGACGAAGATAAGAAGAACCAAGAAAGATTGCAAGATCTGGTGGAGAAGCTTCAAGG
TAAAATCAAAACCTACAAACGCCAGGTGGAAGAGGCGGAAGAAATTGCTGCTGTAAACTTAGCAAAATACAGAAAGATCC
AACAGGAAATCGAAGATTCCGAAGAAAGAGCTGATCAAGCTGAACAAGCGTTGCAGAAGCTGAGGACCAAAAACCGCAGC
TCCGTTTCGACAGCGAGAGGCGGTAGCATGGCTCCCGGCATGACCACAGTGAAATCGACATCACTCTCTGTTTCGGCTTC
GAGCCGACGACAAGGTTCTAGTGTTTCCAGAGAGAGCTAAGTCAGAAATTCTAGTAACACAACGATAAATTATTAGGATT
TACTTAATTGACATTATTTGCATGTTATCAATTATTGGACACAATTTAGTGATGCTTTTATTTAATAAAATTTACATTTT
GAAAAAAAA
>SMED30015552 ID=SMED30015552|Name=POU domain protein|organism=Schmidtea mediterranea sexual|type=transcript|length=1696bp
TTATTTTTAAAGTGTATCTTAATATTTCAAGAAACTGTATTATAAATTTCACAAATTAAAATATTTGGGATATTTGTAAA
TTGTTAATAACTCTTCATAAATGTCTCAATCAGATGGAAATTTCAACTCCTTTAAAAATATTCACATTGAACAGTTATTT
AAAATTATTCAACACCAAGCAGAACAAATTGAAGCACAAAAAGTTCAAATAAACGCTTTAATAAAGTCACAGCAAACGAT
ACTAGAAAGTATGAGACCTCAAAATGATGCAAGCGTCAATTTAAATCCTTTGTCAATGAATTACCTTAATCACAATATCC
GACAATCTTTGTTGAATTCTTCCTTAATGCCTTCAAAAAATATTTTAAACAATCCCTCATTAAATTGTGCTAGCAACAGC
AACAGTAACAGCAGCAACGTCATCATCAACAACAACAACAGCAATATTGATGTTTCAGACCATTTTCTTCAAGACTACAA
TAGTCGATTAACACAACAAATTCTTCTTCAGCCTTCCTTGAATATTCCTCGCTCAAATCCCACAATAAAATCCCCTCATT
TGCCAACGAATATTGGAGGCTCAGGAAATCTAACTCCCACAACTTGTCTTAGCAACGAAACATCTGAATTGAGACCCATG
GAAAATTGGTCCAATTTTAAGTTGCAAGCTCATTCCGATACTCAAAACGTAATCAAAAAAGAAGTAGCAGATCAATCAGA
CAATTGTAACAATTCTGAAATTCCAATTATCGAAAATGATCCAAATGAAGATGAAGGAGAAATGGATTTGTCATCTCTCC
AAGAGTTCGCCGCAAAATTTAAAATGAGTCGTATTGAAATGGGATTCACGCAAAGAGATGTCGGTGTTTGTTTGGGTTTG
ATGCAGGGCAACGATTTTAGCCAGACCACACTGAGTCGGTTTGAAGCTTTAAATCTGAGTTATAGGAATATGAAGAAACT
GCAACCGTTGCTACAAAGGTGGCTGGATGATATTCAACGAGAAGGTCGTGATGCCGTTATGAGTCGCATAACGGCTTTTT
ATCAACAACAAAAAGCGATGGTGGAGGAAGACTGTGATAACAATGACCTCCAAATAGTCAGTGGCTCAAGTAGGAAAAGA
ATTGCAAATGAAGGATTGCTAATATCCGACGAGTTACCCGAAGAACGACATTCAGTCAATGCAGATTCGACTACTATGAA
TTCCAAATTATCACAAAATCAATTACTTCGCACTTGTATAGGAAATATATCGAATAGTCTTTCACCTAGTCTGCTCGGCA
ATTTGACTAATGGGTTTTCTACAAGTCTTTCCACCTCGCTCCCAACTAACCTCACTCACAGTTTTCCGAATATTCAGTTT
CAATCTTCATTACTCCACGACCAAATCGCCAACGGATTCAACAGAAATCTTTCTTCCCAACCAAAAAAACGACGTAAACG
ACCTCTAATATTGGAAGAAGGCAAATCAAGATCGGAATTGTTTCCCGATATTTTGAGACCGTAATCAGACCGGTGGAAAA
GAATCAAAAAAACAACACAAGTTCTACGGCTATTTTATTTATTTTCGCCATTTCTCGCATTGTAATATTTTGTAATTATT
AATATCCTATCTATTGTTTTATAAAATCATATTTTTTTCAAATTGAATTTTTTATTATTTTTCATTTTTATTGACCTGGT
TTGTGAGTGTTCGGAA
>SMED30027428 ID=SMED30027428|Name=forkhead box A-1|organism=Schmidtea mediterranea sexual|type=transcript|length=2149bp
ATCACTTGGTGCTGCTTTTTGCCCCGATAAATCTCTCGGTGCTCCAAAATTTGTGTAAGCGAGCAAAAGATGATTGGGCA
ATTTCTAGCACGTCATTCGGCAACGATGTTTACAAGTTTCTCTGATTGGATGATAAAGTAGAAACTTTCCGAAAATGAAA
CTTCTCCGAACATGCGCATACCGACTGAAAATTAACTCTAAGCCAGCCAATGGAATTGAAGCAAAATTCAAGAGAATATT
AATTGACTTCGCGAATGAACTCAAATTTTCTATTGGCGTATTGTGTGTGCCTCATTGTGAGTGGTTGCGATCGCATCTCA
AATTATTTACACACAAAAAAATAACGCACACAGACATTGAATAGATATATTTGATCAAGTCAATATCACAATGTCAATAT
TAGCTTTTTCTGAAGAAAGAAGACGAACTAGAAGAAAAACAACAACGAAATAGATTCGATAAGGCTACTGAGCGATTTTC
ATAATCTAAATATTATTTTATTATTGATACGGACAAAAATTGGTCTTTTACCAAGTACTACTAGATTGTTGATAAAGAGA
GGTTATTTAGATGCTTGGAAAAAATCCTTATGAAACTGCAATGAGCAACGTGTATTCTCTACCTCCGGGAGGTTCTATTT
ACAATATGAACCCGATGAGTATATCATCAGCTGGCTACAACTCTCAACAAGTATCAACACTATCGTTGAACTTGACCGGA
ATCGGACCTCATTCATTAAGCCCAATGAGTGCAAGCATGTCGGGTATAGCTGCAATGGCCGGTGGAATGAGACAAGGTCT
TGAGTTGGGTCTTGGTAGAAGTGATAGTCCAAGAGATAAAAATTCAATTTCCAATAACAACCGACCATATCAAAGAAGTT
ACACTCATGCCAAGCCTCCATACAGTTATATAAGTTTGATAACAATGGCGATTCAAAATTCTCCAGTAAACATGTGCACT
CTATCGGAGATCTATCAATTCATTATGGATCATTTTCCATACTATCGTCAAAATCAACAGCGATGGCAGAATTCGATTCG
ACATTCTTTGTCCTTCAACGATTGCTTTGTTAAGGTTAGTAGAAGCCCAGAAAAACCAGGTAAAGGCTCATATTGGACCT
TGCATCCTCAATCAGGTAACATGTTTGAAAACGGTTGTTATCTCAGAAGACAAAAGCGATTCAAAGATCCACACAGAGAA
ATCGGCAGACAGAGTCAAAGAGCTGCCACTGGTCCTGGATCAAATGTCACAGAAAACAATCACGACAACGCATCGCAAGA
AGCTAGTGATAACGCAGAAAGTGATACGAAACCCAACATCAAGCAACTTGATTTATCAAGCGATCTCTTAACTAATCAAG
GTCATAATATTAAAAATACTAATCCAACTTCTGTTAGTCAGAGTTGTTCGATGTTTCATCGGAAAAAGGAAAACTGCTCA
CCAGTAGAAATGAAATTGAATAACCAAAACCAACAATCAAACCAGCAAGAACATCCACAAATCCATTACAATCCCAATCA
GCAATTCTACTCAAATCAGCAAAACATTTTCCAACAAAGTTCTCTTGATCATTACAGTCTATTAGCATCCGATGATCCTC
TTGGTCAAGGTATGCACTTGCCACCAGGTGCAAATAGTGTTTTCGGACTTTACGGGGCACATAACTTACCAAACGATGAT
CAAATTTCAGTGTCATTACCATCGATATCCTTATCCGGACATCCGTATGACAATTTATCAACAGCAATGGCATATCAATA
TGAAGCATCTCAACACAATTCTTCATTACTAACGACAAGTAATCCGTTCTCAATAGATCGTTTGATGCATCCAAGACTAG
TCGCTGCAGCGATGGGGGTCAGTCCCCATGATACTCTATACGCAGGAGCTACCGGCCCATCAGTTGATCTAGAACACATG
AAATACTACTCAAACTACAACAATGTGCCTCCTTATTCCTCTGCAATGTCTGACTACTACAAATATGTACAAAATCCTCA
GCCGGGCAACAGCGACATGAGTCTTTGAATTGAGTCCATTGAAGTCTACGGCAGTTTCCTCGAAATTTCACATTCAACCA
GATGTTTATCGCCTAATATAAAGCTGTGTTTTTTTATTTATTTACAATTAAATCTTTGTACAAGAGATC
>SMED30005748 ID=SMED30005748|Name=HNF4|organism=Schmidtea mediterranea sexual|type=transcript|length=3066bp
GAAGAATAACAAAGAAATTTATTGCCAAACTAAAATACCCGGCCTATTTCATCAAAAAAAAAACAATCAAAAACACAAAT
CGCTACGAAAACTTGTATAAAAGATTCACATTAAAAGTCATCAATATTCCTCTTTTTTAAAGACTGATGATTCAGATTCT
TGGATCCGAATGGGGTGATCTGTTAGGAAGTGAGCTGTTGACGAGATAGCTGAATAGTTTGATCTGGAATTGTGCTGTTG
GTTCGCTTCCGATGATCTGTTGGCGATTGATTGAATTTGGCAGTTTTCTATCGGCGAGTTTGGAAGCGACTTGGTATAGG
CCACAAGAAGCGACGGATCAACGACATTTTGAGTGGAATTCACAGAAAGAGCTGGGTGGATTAGGTCATCTGAATTGGAA
GAATTTGAGCAGTTATGATTTGCGTCACTGTTATAGGCGTATTTTGACTGGTTGATCATGGGGAACCCTTGATCAAGGTT
GACCAAGAAATGGCTGGTATCCTGTGATAAATTGGAATGCTGATAAAGATTGATGGAATCATTGACGCATGAAAACAGAG
AAGGAACGCCGACTCTTTCATTTTTCAGCATCATCCCATCGCTGTTGATTATCATACCGAAGTTGCTATTGTCCTGGGAA
TTTTGCTGCATTATTTCAGTTCCTTGATTAACAAAAGCTCGATATATATTTCCTGAAGGCATCGACGTAGAACTTGGCAA
ATATGCACTCCATATTTTTGAGGCCGCTTCAGGTGATATGTTTTGGTTGTTGATATTGAGTAGGACACTGTCTGGCATGA
AAGGGGGAATTCCGTAGCAGTCCTGGTAAGTATATTTCGGTGAGGAAATCCTGTGGGAGTTAATTATATCCATGGAATCG
TTTTGATTATTGTTATTGTTCCCATCTGTTTCTGGATCTATGCTGAATACCCCTGGTGGATTGTCACCCAATAATGTTTC
AGATAACAAGCTATCAATCTCGGCTAATCCTGTCATTTTCATGAACTCGACCTTCTTTACCATCAGCTGGGTGACCAATC
TCAGATCAGGAATTGTCAGCAGCAATTCCCCGAATCTACCCGGCTTGTGGTATTGCTTGTCATTCATGTGGTTCATCAGG
TCCATTTGTATTTGGTATCGGCATCGTCTCACGAACTCCTTGCCAGACTCGCTGACTCTCGGATCAAAAAATACGATCGC
TTTCAAACAAGCGAATTCTGCGTCATCCAATTGCAATTCTCGCAGCGGAATGAACAGGTCGTCAAGGATGTGACTGGCTA
TTTCAGCGAAATGTTTGTCACTGGAGTTCCTACTGATGATAAGATTGTTTCCCAATAATAGCACATCGTCTTCGTCCAAT
TGAAACGACCGCCTTATTACTCCAAGAATTAGAATTTCTCCCGCGTGTGCTTTTAAAAGGGAAATTTGATCGCTCAAGTT
TAACTGTGAAAAACAAGGCAAAGACTTTGCCCAATTCACTAGAAGAAATAACTGATTTTTCATGGACTCGCAAACATCAG
GCACATCGGCATATCTGTTCATGTATTCCTGAGGATTCGGGGGCTTTTGTATTGCAACCCTTTGCTCGGCCTGCATCAAC
TGAGAAATGGAGAGAATGACATTCGGAGGAATATCATCAAACGACGATCTTCGAGTACTGATCCTATCTCGTTCATTTTG
AACCGCTGCTCGCTTCATTCCTACAAGAATGCATTTCTTTAGCCGACAATAACGGCATTGGTTTCGTTTATCCTTGTCCA
TTTTACAATTCCGATTGTACTGGCAAGTGTAGTTATTCTTCTTCCTCACTGACCTCCGAAAAAATCCTTTACAACCGTCA
CACGAGAAAGCCCCATAATGTTTTCCTGTAGCTTTGTCACTGCAAATGAGACAGAGTTGATTGTTTTCCATTCCTTCTGG
TCCCGACGCGTTAGACGCATAGACTTCATGTTGTTGCGAGGACACTGAAACGAAAACTATTTGATAATATTTGTACAAAA
CAACTGACAGATGGTTTCGGAAAAGGTTCAATGTTCAGCCATAGTTCAGCGAAAGGTCAAATTCCTGAACTACTATTTCA
AGGTATAAATTTGAATATATTGGTTTTGCCTCGTGCTAACAAAGTCCAGAAATATAATAAGCAAAGTCTGAATCGTGTAC
TTTGGATTTCTAGCCAGGGGCACACAATAACCTATATTTAATAGCAGTTCATTGCGTTCACTCGTTATTTCTCCGACTGA
ACTGCATTGCAGGAGGATCCGCAACCATGGAGATTTCATAATTATTATCAAAATGATAATAATAATAATTATTATGAGGT
ATTGTAAAAGAAAACGTAACTCACTTTCACTTTCCAATGGATATTGCATCGAGGAAGCGTTGCTGGAAATGGAAATCGGA
GCTGAATTGCTAGTCAATAGTTGATAGGAGAATGCTGGATTGAGGTTCGGATGCTGGGTAGACGTCATATAGAATTGGCT
CGACGGATTCGGTAAAATTATGGTGAAGTTTACAATCGAAGCTCAGATATAACGTATTATAATGAAGTATTATTAATAAA
CTGCAAGTAACGTGCGCGCATTCTGAAGTGGTTGATTGAGAATAATCAGTTCTTATTCGACAAATTCAGATAAAAAAGGC
ACCACTCGAATTGTTCAGCTGCGTCATGTGAATGCCGGAACGTTGTGCGTTGGCTGTTGGCAGATCAGCGCTCCGTGTTC
ACTGCAATCCGGATACACCAGACCACTTGAAATCGTTGCGAATTCAGCGTGTCGAGTGCATTGTGCGAGATCAGTATCGG
TGTGTATGTGTGTCTGCCTGTGTGCGTGTGGATGTGTGTGCTTGTCGAGTCGGTTGTGTGTTTATGAGTGTAACTGTGTC
GATTTGGTCGAATGCAGTAGCAGATAGCTGAAATGCGATTGTTCCTATTTTTCATTTAATATTGTTTTATGTTTTGTTTC
TATTTTTATAATTTAATTTATCAGTTTTTGTTTAGATTATGCAATCATTGTTTAATTTATTTTTGTCAATTTAAAATTTA
TTATTAATTAAATTATTATAATTATT
>SMED30014019 ID=SMED30014019|Name=Intermediate filament protein|organism=Schmidtea mediterranea sexual|type=transcript|length=768bp
AATTAGAAATCGCAGCCTATGCAAGGTTATTAGAAATTGAGGAAGAGCGAGTTTTAAGTTCCGCTGATCAACGTTCTGTG
ATGTCTCAATTTATGAGTTTTTCAGCTGGTGGTGGAGCTGGAGGTTCTAGTGGAGTTGGAGGATCAGGAGTTGGAGGATC
AGGATTTGGAGGATCTAGCTTTGGAGTGTCAGGCACCGGACGATCAGGCTTTGGAGTGTCAGGTACCGGAGGATCTATGG
GAGGAAGTAGATTTAGTGGAAGTCTAGGACTTGGGGGCGGGATTCATTCAAAAAAGATGTCTTCAAGAAGAACGACAACG
GGAAACATTTCTATTGCCGAAAGTGATGAATCTGGGTTATTTATTGTTTTAGAAAATATGGGAAGTCAGACTCAAAGTCT
ATCTGGCATGATGATTAAAAGAAACATAGATGGCGTAGCAAACTTTGTGACCTACTCAATTCCGAAGAAGATCGATCTAT
CCGCTGGTGAAAGAATAACAATTTGTGGACATGGAAAGAAAACAGGTAAAGGAAAAGAACTAGAAATAGAAGAAGATTCT
TGGGGATTTGGAGAAAACGTCATTACTGCGTTATGTGATGCGAATGGCGAGGAAAAAGCGGCTCTTTATCAAGAAATAGT
GGCTTAAATGGCATAGCCGAAAGACTTGCAAATTTAACCTTTTATTTAATGATTTCATTTAATGTTCCTCGCGTTCATTA
TTTGTTTGCTTCTTTCTAGATTAAAGTTTTAAATTTGTTGAAAAAAAA
>SMED30008639 ID=SMED30008639|Name=MyoD-like protein|organism=Schmidtea mediterranea sexual|type=transcript|length=1606bp
GTTCATATTACCAACTGTTCGCCATTAATTCAGCCTGTTTTACGCCTCGTAACAACTGTTACTTCAACAATACCGATCCA
GCCCAATAAACAATGATACACTCCGGATCATTTCCAGGTTCCACTTGTCCGATTAACTATGACTATTCCGGTCCATACTC
AGCAGGTCTTGAGGCAGCTCCATTTTTAGTCAATCAAACCTGTCCATATAACACTGGAACAAATACAATGCCTCCTTCGT
TGTCACTAGTAGATAATATGATGATGAATAATTCCATGCATCTTTATGATAGACTGAATACTCATCATTCAGCGTCGTAT
TTTCCTCAGATTTTAAACAAAATCGAACAGAAATTACCAACAACCTCGAGTAAAACCACTCCATCAACATTGGTTTCAAC
ATCCTCCAATTCCATATCGAATTCTGAGTTTAATTCGTCGCGATCTTCGTCACAATTTCCAACATACGATTTGATTTCGT
CACTCAATAATAGCAACTACGACATGTACGATTCGCCCTATGCCGCCTATCTGGCCGCTGGTTATTATCAATCTCCTGGA
TTTGCGCTCAAACAGTACGACGCTGCTGGAACATCTCAACCATCAATGACAGATTTGTACCATTCATCACTAGCAGCCAG
TTCAGGCTTAAATTTGAATTATTCACAGCATAAATCGAGACATTTATCAGAAGATGCTGGATCATGCCAAGATCCGCAGT
CTCAAAATATCACAAGCAATAATCATCATAACAATTCGTATAGTCCAACAACTGCTGATTTCGGATCACCAAAAGCCTTA
TCAAGTAGCAGCCAGTCCATAAATGATGGAAGACAACCGACCAGTAGTAATTCCAGTCCTCAAAAACTATCACTAGCAAT
TGAACAACACTTATGTCCAAATAAAGAGGAACATGTGCTAGTTCCAGGGTCACATGGTCAATGCTTGTTGTGGGCCTGTA
AAGCTTGCAAAAAGAAAACGGTTCACGTTGATCGCCGCAAAGCGGCAACAATGCGTGAAAGACGTCGATTGAGAAAAGTG
AATGAAGCTTTTGAAACATTGAAACGAAGAACGAGCGGTAACCCAAATCAAAGAATGCCAAAAGTCGAAATTCTAAAGAA
CGCGATTGATTACATTGAGAGTTTGGAAGAGTTGTTGAATTCCGTGGGAAAATTACCTGCAACCGTTGGAGAATTGAAAG
AACGCCGCAAAAACAACCCAGAAAATTCGAGAACAGCAATCGCCGGAAAACAAAAGCCACTAAATAAAATTAATGATAAA
CAGAAAGCTGAAAACAAATATCCCGAGACCATCTGCAAACTCCCGACGACTAGGAAATCCTATGTTGCTTCCGACCTTGA
TCAGCAGATTTATCAATCAAACATGTTGAAGAGTTCAACTCTTGATCAACTTTCCTCGATAGTAGACAAAATCTCTACGA
CTAATACAACAATTGCCGACCATTCAATGGACGCTAAGCCCGAAAATAGTTGAATACATAACAAAACGCAACAAATATTA
TATTAACCTATTAATTTTGTTCTAAACTCGTTTGCCCTGATACATATTGTACATTTTCTTTCAATAGAATGTTCTTATAA
TTATAA
>SMED30035904 ID=SMED30035904|Name=PROG-1|organism=Schmidtea mediterranea sexual|type=transcript|length=728bp
GCAATATTTTGTCATGAATGTTTAATTGAAAAATATGAAAGTTTTAATTATATTTCTATTTGTGATTGCGTTCGCGTATA
TTGAATGCCGTCCTCAACATCTATTGGGGTTCATGAGGAAAGAAAAGTCCAAGCTAGAAAAAGACTCAGCAGATGACGTG
AAACAAGGAAAAGATTCGGATAAAAGCTCTGATAAATATTCACATGAAGGATCTAATAAAGAGTCTGACAAAGAGACAGA
TAAAGAAGAAGAAACTAATTCTAAGACTTCTGCAAAGTCTCCCGCCAAATCAAGTAGACGAAAGAAACAAGTGGAAAAAT
TGGAAGAAGAGGAAGGAGAAACTTTAAAAGAGAAATTAAAAGGACACATTACGAAAGCAGATTGCATGAAATTATACGAA
GAAACTGATAAATACTGCGATCCACTTTATTTGAAGCTCGACAAAGATTGCGACTGCAGATATGTGAAAGCATTGAATGA
CTACACGAAAAGATGGGACAGTTATCATTCCAAATGTGGTAGCAAATATGACAAAAAATGCCGAACCGAATTGCGAGCTT
TGGAGAATTGGTTTAATGGCCTTTATGATTCAATCGACGGCCACTGTGACGCAGAATATGACGCGCTGGATAAATATTGT
GACGTAGAATACAACAAAGTTGATCAATGTGAAAAAGGACAGAAAGTGTAATAAAATTGATTTGTAATAAAATTTATTAA
ATAAAAAA
>SMED30032169 ID=SMED30032169|Name=Glycine amidinotransferase, mitochondrial|organism=Schmidtea mediterranea sexual|type=transcript|length=1380bp
CGTGTTTCTGCCAGAATCCAAGTTTGGTAGGGTGGGTGGAGTAGGCTTTTATATCATATCTAAATCTGAAAATGAATTTT
TATCATCATATTTTCTCAAATGTTGTCTGTATTACGACAAATTAGCCATTCAAAATCTGCCATTTCTTCGAAAATATCGG
TAATTCACATTTCAAAGTACTTCGGTCATTATTCATGCCACGATGAAAAGAAACAGACTTCTCCAGTTTGGGCTTGGAAT
GAATGGGATCCACTTGAAGAAGTTGTGGTCGGTGTTCCTAATAATGCAACTTTGCCCTTTCCATCACAGGAAGTAGCAGC
CACTTTTGGACCCCCTAGACTTCCATTTTTTCAAAAACATGGTGGCAAAAAATTCTCAGAGGTTATGGGTCAAGAAATGT
GGATGAAAATTGAAAATGAAATAAAAGAATTAGTAAGAGTATTAGAGGGAGAAGGTGTAAAAGTTGTGCAACCAGAGGCT
ATAGATTACGATCAAAAGTGGACTACGCCAAAGTTTTCTTCAACAGGTTTGTATGCCGCTATGCCAAGAGATTTTTTACT
TATCGTTGGTGATGAAATCATTGAATCCCCTATGGCTTGGAGGTGTAGATACTTTGAATACGAAGCATACAGACCACTTA
TTCGAAAATATTTTAATGAAGGAGCCAGATGGACATCGGCTCCTAAATGCACTATGGAAGACGCTCTCTATGATATGGAA
TTCGATGTTGCTCAAGAGAAACCATATGTGTATGTAACAACTGAATTAGAACCATGTTTTGATGCGGCTGACTTTACAAG
AATTGGCAGGGATATTTTTGGTCAAAGAAGTCATGTAACCAACAAAAAGGCAATACAGTGGCTCCGGCGTCATTTAGCTC
CTAGTGGAGTCAGGGTCCATGACATTGAATTTGACTCTCTCTATTCAATGCATATTGATACCAATTTTTTCCCCGTGAAA
CCTGGTATGTTTTTACTTAAAGATGGACTCAATTGTAAAACCCCGGATCTATTAAAAAAATCTAAATGGGACATAGTCAT
TATGCCAAAAGGACCTCATCGAGACGAGCATTACAAATGGATGAATTATCACGGATTACCGAACAATATGCTCAGTATAG
ACACAGAGAGGGTCCTAGTTGAGAAAACCGAAGAATACATGATAAAATTCATGCAAGAAATTGGATTCAAACCGATTCCA
GTTTCGCTTAAATTCGCAAATAATCTAGGAGGGGGCTTCCACTGCTGGACAGCGGACATCAGACGGCGAGGAAAACTAGA
AAGCTATTTTGAGTGGCCAGATGAGATTTTGGAGCGATTGACTCCATAAATCGGAAATATTAAATTAAAAAATATTTTTT
GTTTTAATTTTCTATACAAT
>SMED30030379 ID=SMED30030379|Name=POU domain protein|organism=Schmidtea mediterranea sexual|type=transcript|length=1625bp
AATTAGCTATGCGAGCTGCTTTATCAGGCATGACTGTGATAAAACTGTCATTATTTATGCGCATAATTTGCCGTTAGTAT
TTGCTCAACCAATTTGACGTTCAACAACAACAGCAACAACATCCACCACCTCCACCAACATCCTCATTATCATTATTATT
ATTGTCATGTTAACAAGCGGACTTAATCAGGAACAATCATCGCCTTTAACAATCGTAGGCATGAGTGAGTCTCAGTTTTT
CAACGAATCTATCAGATATTTGCCTCCGGAAACCGTGGAGAGACAAAGCCAATTTTATGCAAATAATAATTATCCCCTTG
GTGAGTTTTGGAATCCACAATCCGAATATGCACAATACTTGCAAAATACCCAGCCAGTCTCGGGGCCTCAATTCATTACT
TCGCCGGCGGTTGGAGAGGCATTCTATCCATCTCTTTATATCAATACTATCCATGAAAATGTATTACTACCTTCGGTTGA
TTCCTTCAATCCTCCCAGTGGATTTATGATTTCGGGATATTCACAGCAAGACCAAACCTTTCCCACGGTCAATGAACCGA
CGTATGACAATATCCCAAACAATTGGAATTATGTTGATCATTGGGTTGATTCCAAGCCAACAACAGTGAATTCTCAGCTG
TTGCCAACCTCGTCAATTTCCGCCGTCCCCATTCAGAATTCGTTTTGGGACTGCGAGTCGATTGACCAATCAGAATATTG
TCCCACAAATGTTGATACCGAGCTCCAAGATCAAATTGATTCCAACGGAGAAATCAGTAGACTCAATCAAAAGATTTCTA
CTGACGAGTTGGAGCAGTTTGCGAAAATTTTCAAACAGCGAAGAATCAAACTCGGATTTACACAAGCCGACGTCGGGTTA
GCCCTCGGTAATTTATACGGCAGCGTGTTCAGTCAAACAACAATTTGCCGGTTTGAGGCACTGCAGCTCAGTTTTAAAAA
CATGTGCAAACTTCGACCTCTGTTAGGGAAGTGGTTGCAAGAAGCCGACTCAACCGGATGTGCTCCGTCCGGTATTGACA
AAGTCACAGCTGCTGGACGAAAACGAAAAAAGCGAACCAGCATTGAGGTTGGAGTGAAAAATGCACTGGAATCTCACTTT
TCTCGACAAAGCAAACCGACAGCCCAAATCATTACACAGCTTGCAGACAGTCTCGGATTGGAAAAAGAAGTTGTCAGGGT
ATGGTTCTGTAATCGGCGACAGAAGGAAAAACGAATGACGCCTCCGCCACTGAATGGAACAAGCTCAGATTTATTTCCAC
ATCCTGGTTTAACTGTCGGAACTGATAGTTCTCCTCATTCCTCATCTCACAGTAATCATGATTCATTCTCGTCCATTGGT
AACCAGTCAAGCTTCAATCGAATAGGTGATCCCTCCTCGACTCATCCCGGAGTCATTGTCACATCTTGCGGTTATTACTA
TGGTCATGTTCCAAATGAATCACTTCTCAATTGTGGGCCTTACCCTCACAACATCATCAACAATTACGAATTCAGACAGT
CCAATGATCCCAATATCCTCAATGTCAAATCCGAGTGCATCAGCTGACTCATGTCGAACTCACCGAAAAGAAAGCTAATA
ATATTCTATTTCAATAATAATAAAA